The human metabolome and machine learning improves predictions of the post-mortem interval
Statistics and reproducibility
The study was approved by the Swedish Ethical Review Authority (Dnr 2019-04530 and Dnr 2025-0249-02). Due to the retrospective nature of the study, the need of informed consent was waived by Swedish Ethical Review Authority. All methods were carried out in accordance with relevant guidelines and regulations, and all samples were irreversibly anonymized prior to analysis. Autopsy cases were obtained from the Swedish National Board of Forensic Medicine. After transportation to the autopsy site, bodies were stored in controlled indoor environments, typically in morgue refrigeration units, which substantially reduces variability in decomposition rate compared to outdoor or uncontrolled conditions. For more details, see Supplementary Methods 1.
For the original study, we included all autopsy cases admitted between 2017-09-01 and 2019-03-14. For the independent test data, cases were collected between 2021-01-01 and 2021-12-31. The inclusion criteria were as follows: availability of femoral blood, age ≥18 years, and completion of toxicological screening using high-resolution mass spectrometry. For more experimental details, see Supplementary Methods 1. No statistical method was used to predetermine sample size. No data were excluded from the analyses; The experiments were not randomized; The Investigators were not blinded to allocation during experiments and outcome assessment.
From the original study period (2017-09-01 to 2019-03-14), all autopsy cases with a certain or probable death date (as described below) were included (in total 4876 autopsy cases). The most frequent causes of death in the studied population, each with more than 100 documented cases, included complications of cardiovascular disease (n = 748), acute poisoning with one or more drugs (n = 659), hanging (n = 572), alcohol poisoning (221), drowning (194), trauma resulting in multiple internal and external injuries (150), and gunshot wounds (119). The demographic characteristics of the selected cohort can be summarized as follows: age, median = 56 years (interquartile range = 39–69); sex, male = 3544 (72.3%); BMI, median = 25.7 (interquartile range = 22.5–29.6); PMI, median = 5 days (interquartile range 4-7).
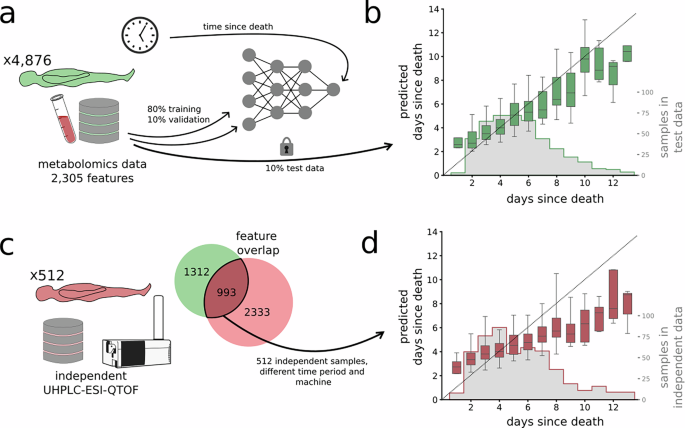
For the independent test data (2021-01-01 and 2021-12-31), we randomly selected 512 autopsy cases with a certain or probable death date (as described below).
Calculating the post-mortem interval
In the database of the National Board of Forensic Medicine in Sweden, the date of death can be ascribed and coded in two different ways: certain and uncertain. For the uncertain cases, a probable date of death is indicated together with the date the deceased was last seen alive. The present study includes all cases in which the date of death is certain (n = 3954). Additionally, cases with a probable date of death are included if the date when the person was last seen alive is the day before the body was found (n = 922). This approach allows for the inclusion of cases where, for example, a person was last seen alive in the evening and found dead the following morning. However, this also gives an uncertainty of up to 48 hours in extreme cases (body last seen day 1 at 00:01, body found day 2 at 23:59).
In some cases, the date of death was ascribed as probable even if the date last seen alive was the same date as when the deceased was found dead. In Swedish forensic practice this is sometimes done to indicate that the death was unwitnessed (i.e., last seen alive in the morning and found dead in the evening).
In the present study, PMI is defined as the time (in days) between date of death, certain or probable as described above, and the date of the autopsy in which sampling was performed. Autopsies are typically performed during morning office hours in Sweden. Therefore, e.g. PMI = 2 can for the cases with a probable death date represent between 32 hours (body last seen and found 23:59 day 1, autopsy in the morning day 3) and 80 hours (body found day 2, last seen day 1 at 00.01, autopsy in the morning day 4).
Software implementation
The post-mortem metabolomic data from the samples were pre-processed in R using the xcms package and CAMERA package, as in ref. 33. This workflow included peak detection, retention time alignment, and feature grouping, and gap filling using the XCMS fillPeaks algorithm, resulting in 2305 metabolomic features. Missing values occurred when a metabolite peak was not detected in a given sample during XCMS processing; these missing peak intensities were imputed as zero values, reflecting metabolite abundance below limit of detection.
The computational analyses were performed in Python v. 3.12.4, mainly relying on the packages Keras v. 3.3.3, scikit-learn v. 1.5.0, numpy v. 1.26.4, scipy v. 1.13.1, matplotlib v. 3.8.4, and seaborn v. 0.13.2.
Data processing
We normalized the data using a log-transform (Eq. 1).
$${x}_{norm}={{\rm{ln}}}(x+1)$$
(1)
where x is the peak intensity and xnorm is the log-transformed expression. The log transformation was applied to stabilize variance and reduce skewness, thereby mitigating heteroskedasticity inherent in the raw data. Following the log-transformation, we standardized the values of each metabolite using a z-transform (subtracting the mean and dividing by the standard deviation of each log-transformed metabolomic profile). This step ensured that all metabolites were centered at zero and scaled to unit variance, facilitating comparability across variables in downstream machine learning models. The log transformation and z-transformation addressed different aspects of the data distribution: the former reducing skewness and heteroskedasticity, and the latter ensuring comparability of variables by placing them on a common scale. Exploratory analyses of metadata suggested minimal batch effects or other sample-to-sample variation, and therefore no additional correction was applied.
We normalized and standardized the dataset prior to splitting. Given the large sample size, the differences in distribution between the full dataset and the training subset are minimal, and the impact on model performance is expected to be negligible. We randomly divided the dataset into training (80%), validation (10%), and test (10%) using probabilistic assignment.
Data harmonization between the different datasets
To harmonize the variables of the independent test data to that of the original mass spectrometry dataset, we aligned the features between the original and new datasets using a peak mapping approach. As such, each feature in the original dataset, defined by its median mass-to-charge ratio (m/z) and retention time (RT), was matched to candidate features in the new dataset that fell within the corresponding m/z and RT bounds. Among these candidates, the feature with the minimal squared distance from the original feature’s m/z and RT was selected as the best match. If there was no such candidate, the feature was set to 0. This mapping allowed the new dataset to be reordered to match the feature structure expected by the model.
Neural network design and training
To predict PMI, we implemented a feed forward neural network regression model using the Keras package. To determine the optimal design of the model, we initiated a hyper-parameter optimization to select the number of hidden layers, the number of hidden nodes in each layer, and the dropout rate for each layer. Additionally, we implemented the option to train the model with a custom attention layer as an input layer such that each metabolite would be individually passed to a single node. The rationale behind this was that such an attention layer would serve as a data-driven feature selection algorithm.
We trained each hyper-parameter setting three times and selected the best performance using an early stopping algorithm with a patience of 25 epochs with respect to prediction error on the validation data. The early stopping algorithm was also used to restore the best weights during training. All tested hyperparameter sets can be found in the Supplementary Data 5.
Training of alternative machine learning models
We trained a compendium of alternative machine learning regression models using the same training data as for the FFNN model. These models included the two linear regression models, Ridge and LASSO, where the respective L2 and L1 penalty weights were selected using the Scikit-Learn built-in RidgeCV and LassoCV implementations. We also implemented a gradient boosting and a random forest regression model, each with the default 100 estimators, as per the default setting on the Scikit-Learn package in Python. Furthermore, we also implemented a K-nearest neighbor (K-NN) regressor and a support vector regression (SVR), both as implemented in Scikit-Learn. We trained the models using the assigned training data, and, for consistency, tested them on the test data. The used settings of the alternative machine learning methods can be found in the Supplementary Data 5.
Extracting learned model structures and feature identification
We sought to extract which metabolomic features were used in the decision-making processes of the respective models. While such an extraction of input-output dependencies is non-trivial for neural networks34,35, we utilized the attention layer of the FFNN to estimate the usage of input variables. By analyzing these input node activations and correlating them with PMI, we generated a list of metabolomic features ranked by importance.
All metabolomics features included in the FFNN were uploaded to MetaboAnalyst (version 6.0) for functional analysis, which is suitable for untargeted metabolomics data, relying on the assumption that putative annotation at the compound level can collectively predict group level functional changes defined by set of pathways of metabolites36.
Metabolomic features significant for decision-making in the FFNN were putatively identified and annotated by reviewing the compound hits from the functional analysis in MetaboAnalyst, and/or by matching the mass-to-charge ratio (m/z; ± 5 ppm) to the online human metabolomic database (HMDB; https://hmdb.ca).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link








