Multi-task genomic prediction using gated residual variable selection neural networks | BMC Bioinformatics

This section describes the methodology for genomic prediction and variable selection using the deep learning-based GRVSNN framework. Our approach is designed to efficiently handle high-dimensional genomic data by improving feature selection and pedigree transformation, leveraging multi-task learning for enhanced predictive performance and interpretability across multiple traits.
Reduced-rank transformation of the polygenic pedigree
Statistical modeling with genetic covariance matrices is essential to understand the genetic architecture of complex traits [51]. Traditional estimation methods often fail when the number of traits exceeds the number of individuals, leading to unstable estimates, noise, and spurious correlations that obscure true genetic relationships [52]. Reduced-rank estimation provides a robust alternative by stabilizing estimates, reducing noise, and preserving key genetic components while maintaining a parsimonious structure [53]. This reduced-rank approach is particularly useful in applications such as animal and plant breeding, where optimizing multiple traits simultaneously requires reliable genetic correlation estimates [7].
The pedigree matrix A is first decomposed using a traditional eigendecomposition, \( A=Q\Lambda Q^{-1} \) where Q denotes the eigenvectors and \( \Lambda \) contains the eigenvalues. The loadings L are then calculated as:
$$\begin{aligned} L = Q \times \sqrt{\Lambda }. \end{aligned}$$
(1)
After that, the eigenvalues are sorted in descending order and used in a scree plot to determine where the eigenvalues are no longer decreasing [54]. This rank value is the reduced rank m that is used to obtain the reduced rank loadings matrix \( L_r \). We integrate these reduced-rank loadings into the genomic prediction framework by combining them with the genomic markers M into a joint tabular input matrix \( X=[L_r,M] \).
Gated residual block
The GR block consists of multiple layers designed to facilitate feature transformation while it effectively controls information flow. The input features, \( X \in {\mathbb {R}}^{P} \), where P is the total number of features, are one by one first submitted to a non-linear layer:
$$\begin{aligned} f_1 = \sigma _{f}(W_{1} x + b_{1}), \end{aligned}$$
(2)
where \(W_{1}\) is a learnable weight matrix, \( b_{1} \) is the bias term and \( \sigma _{f} \) is the exponential linear unit (ELU) activation function. The ELU function with \( \gamma > 0 \) is defined as:
$$\begin{aligned} \text{ELU}(x) = {\left\{ \begin{array}{ll} x, & \text{if } x>0 \\ \gamma (e^{x} – 1), & \text{if } x \le 0 \end{array}\right. }, \end{aligned}$$
(3)
where \( \gamma \) is a positive constant that controls the saturation point for negative values. We use the default value \( \gamma =1 \). Unlike the ReLU active function, which set all negative inputs to zero, ELU allows small negative outputs, which is helpful to push the mean activations closer to zero, improving convergence speed and reducing bias shift [55]. Following the non-linear transformation, a linear layer with dropout is applied to prevent overfitting that yields an output \( f_2 \), which is connected to a gated linear unit:
$$\begin{aligned} h = \sigma _{h}(W_{2} f_2 + b_{2}), \end{aligned}$$
(4)
where \(W_{2}\) is another learnable weight matrix, \( b_{2} \) is the bias term and \( \sigma _{h} \) is the linear activation function. To regulate information flow, a gating mechanism is then introduced as:
$$\begin{aligned} g = \sigma _{g}(W_{3} f_2 + b_{3}), \end{aligned}$$
(5)
where \(W_{3}\) is another learnable weight matrix, \( b_{3} \) is the bias term, and \( \sigma _{g} \) is the sigmoid (or hard sigmoid) activation function. The gating vector \( g \in (0, 1)^{n_{h}} \) determines how much of the transformed features contribute to the final output. A residual connection is then incorporated to stabilize training and retain original input information as:
$$\begin{aligned} z = g \odot h + (1 – g) \odot x, \end{aligned}$$
(6)
where \( \odot \) represents element-wise multiplication. This operation allows the model to learn which input features should be set to zero and therefore be canceled out. The final output is normalized using layer normalization as:
$$\begin{aligned} {\hat{z}} = \text{LayerNorm}(z). \end{aligned}$$
(7)
Note that layer normalization ensures stable training by normalizing activations across feature dimensions, mitigating internal covariate shift. The output \( {\hat{z}} \) is then passed to subsequent shared layers for further processing. The GR blocks are employed locally on each input but share information over inputs and tasks via the subsequent VS block and multitask loss function.
Variable selection block
The VS block is used to identify the most important features globally across all inputs. This addresses the challenge of selecting a subset of informative genomic and pedigree loading inputs that contribute to the phenotypes. The importance of each transformed feature \( {\hat{z}}_{i} \) is determined via a softmax function:
$$\begin{aligned} w_i = \frac{e^{{\hat{z}}_{i}}}{\sum _j e^{{\hat{z}}_{j}}}, \end{aligned}$$
(8)
where \( w_{i} \) represents the relative importance of feature \( {\hat{z}}_{i} \). The softmax function ensures that the weights are non-negative and sum to one, allowing for a probabilistic interpretation. The selected feature subset is then obtained by re-weighting the input features as:
$$\begin{aligned} z^{‘}_{i} = w_{i} \cdot {\hat{z}}_{i}. \end{aligned}$$
(9)
This operation can adaptively scale the features, enhancing the influence of more relevant ones while suppressing others. Features with higher weights are retained with greater influence, while those with lower weights are down-weighted. The transformed features \( z^{‘}_{i} \) are then used to obtain a multi-task prediction \( \hat{{\textbf{Y}}} \) for N individuals and and T traits. The model is trained by minimizing the joint multivariate MSE loss function over all traits:
$$\begin{aligned} {\mathcal {L}}_{\text{MSE}} = \left\| {\textbf{Y}} – \hat{{\textbf{Y}}} \right\| _{2}^{2}. \end{aligned}$$
(10)
By combining the GR and VS blocks, the proposed framework performs both local feature-specific and global variable selection. This approach parallels the functionality of global–local shrinkage priors commonly used in linear Bayesian variable selection methods [46]. The gated residual (GR) block combines residual connections, which preserve linear additive genetic effects, with nonlinear transformation layers that enable the model to capture non-additive genetic effects. Hence, the GR block extends a multi-layer perceptron (MLP), where activation functions handle non-linear dominance effects and hidden layer nodes model epistatic interactions between genomic markers. The variable selection (VS) block operates globally, reweighting features across the input space to prioritize the most informative additive and non-additive contributions. Moreover, the residual connections and gating mechanisms enable the model to focus on the important inputs, avoiding the huge number of variables typically encountered when modeling epistatic effects with linear models. Finally, the output layer integrates the selected features to produce multi-trait predictions. By distributing additive and non-additive modeling responsibilities across these distinct components, the GRVSNN framework effectively balances interpretability with predictive flexibility.
Training procedure
To effectively train the GRVSNN model across different datasets, we begin by preprocessing the pedigree matrix A using dimensionality reduction. This is performed by calculation of loadings, where eigenvalues are computed, sorted, and truncated at the elbow point to retain the most informative components. Specifically, we extracted 169 components for the mice data, 797 for the pig data, and 138 for the loblolly pine data (see Fig. 2).
Then the GRVSNN model is trained using an optimization strategy that balances predictive accuracy and computational efficiency. Several hyperparameters are fine-tuned using Bayesian optimization (BO) [56], including batch size (64–512), learning rate (0.00001–0.001), and dropout rate (0.1–0.5). To prevent overfitting, early stopping is applied, terminating training if validation performance does not improve for 25 consecutive epochs. Additionally, efficiency of the BO is enhanced by dynamically monitoring the test loss: if the test loss remains stable for 30 consecutive iterations, the BO is stopped to avoid unnecessary computations while ensuring convergence. To improve regularization and sequential dependencies, we integrate the GR block with \( L_2 \) normalization, applied to ELU and sigmoid activations. We also evaluate the hard-sigmoid function to accelerate convergence and enhance sparsity. The hard-sigmoid function is defined as \( \text{max}(0,\text{min}(1, 0.2\alpha _{i} + 0.5)) \), which can ensure that values are smoothly mapped between 0 and 1 while maintaining computational efficiency and preventing vanishing gradients [57]. We also explore the sparsemax activation function as an alternative to softmax for feature selection due to its ability to enforce harder sparsity. However, empirical results showed no significant improvement over softmax in our multi-task genomic prediction tasks. Thus, we opted to retain softmax for its stable performance and easy interpretation. The model is optimized using the Adam optimizer with weight decay for additional regularization. Training is performed using 5-fold cross-validation to ensure robust performance across datasets. Specifically, the dataset is randomly partitioned into five equal folds with each fold serving once as test set and the remaining four as training set. To obtain reliable performance estimates, we repeat the 5-fold cross-validation procedure ten times using different random seeds, resulting in a total of 50 cross-validation runs per method. The final performance metrics are reported as the mean ± standard deviation across all repetitions, providing an estimate of the variability due to random partitioning. Random splits are generated using stratified sampling to preserve the distribution of trait values across folds. These combined strategies enable effective feature selection, improved generalization, and enhanced prediction accuracy in high-dimensional data. For feature selection, a threshold of 0.05 is applied to the feature importance scores \( z^{‘} \) in equation (9), with the number of selected features determined by counting all features whose importance values meet or exceed this predefined threshold.
Other settings
LassoNet is implemented with two hidden layers, and its key hyperparameters – hierarchy coefficient M (range: 10–50) and regularization strength \( \lambda \) (range: 0.0001–100) – are tuned using Bayesian Optimization (BO) over a maximum of 200 iterations. Additional parameters, such as batch size and number of nodes per layer, are also optimized via BO. A 5-fold cross-validation approach is applied to select the best parameters based on the validation MSE. The BO tuning process applies early stopping which means that it continues until a predefined stopping criterion (\( 1e^{-5} \)) is reached. To further enhance parameters optimization, we leverage the tree-structured parzen estimator (TPE), which ensures diverse candidate suggestions across iterations while incorporating new recommendations from BO [56].
In addition to LassoNet, we also compare with three widely used traditional Bayesian regression methods: Bayesian Lasso (BLasso) [40], Bayesian ridge regression (BRR) [41], and BayesC\( \pi \) [42]. BLasso assumes that marker effects follow a double-exponential (Laplace) prior, which induces a strong shrinkage toward zero for small effects that encourages sparsity. BRR assumes Gaussian priors on marker effects, leading to homogeneous shrinkage across all markers without variable selection. BayesC\( \pi \) performs variable selection using a spike-and-slab mixture prior where a proportion \( \pi \) of marker effects is assumed to be exactly zero while the remaining effects \( (1 -\pi ) \) follow a Gaussian distribution. These different priors result in different shrinkage and selection behaviors, which we benchmark against the GRVSNN framework. For these Bayesian models, test predictions are obtained via Gibbs sampling with 6000 iterations, discarding the first 2000 iterations as burn-in. Variable selection for the Bayesian methods is performed by checking if the \( 95\% \) credible intervals overlap zero or not. Implementation is carried out using the BGLR package in R using default parameters [58]. To ensure numerical stability and efficient convergence, all response and explanatory features are centered to zero mean and variance one before optimization, eliminating the need for an intercept term. For performance assessment, we compute the average test MSE, Pearson correlation coefficient (r) and distance correlation coefficient (dCor) [45] for all traits of each dataset.
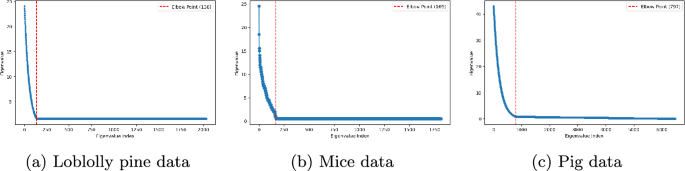
Sorted eigenvalues from the pedigree decompositions for the loblolly pine, mice and pig data. The dotted red lines indicate where the reduced rank selection were performed
link








