Interpretable X-ray diffraction spectra analysis using confidence evaluated deep learning enhanced by template element replacement
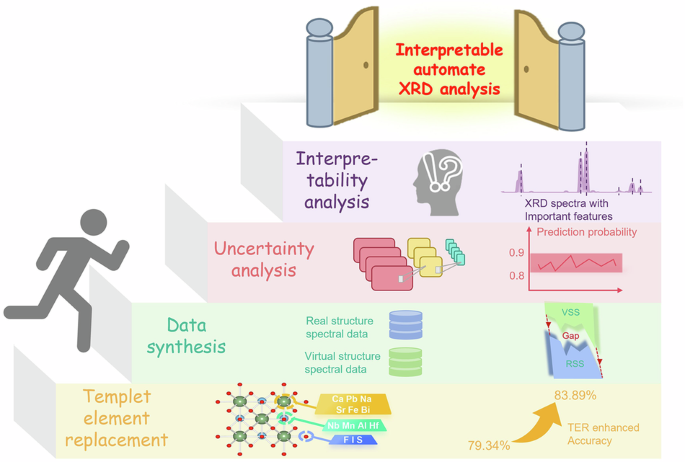
Framework for XRD prediction and its analysis
VSS are simulated within structural templates, covering a wide range of hypothetical and potentially physically unstable structures. This broad diversity supports the learning of general structure-spectrum relationships, but may also introduce significant discrepancies from real experimental spectra. In contrast, RSS are derived from experimentally confirmed, physically stable structures in the ICSD database. In this study, we consider that their spectra more closely resemble real experimental conditions compared to VSS, and thus serve as the ground truth for model evaluation. To effectively bridge the physical discrepancies between real and virtual structure spectral data, data synthesis was performed on both datasets according to their classification labels. By structuring the data in this way, we can directly test whether the mappings learned on VSS enhance the model’s predictive performance on real structure XRD spectra, thereby improving phase identification accuracy on experimental spectra.
The architecture of the B-VGGNet, as presented in the paper, was adapted from the VGGnet used in computer vision. This adaptation was employed to classify and predict crystal spatial clusters and structures. A Bayesian layer was added to the penultimate layer of the network to evaluate model uncertainty. The Bayesian layer utilized three methods: MCD, VI, and LA. Ultimately, the degree of contribution of each feature to the classification was calculated, aligning with its physical significance. The model’s classification results were then analyzed to enhance interpretability. This approach integrated the power of deep learning with Bayesian statistical methods, providing a comprehensive tool for the analysis and interpretation of XRD spectra.
Dataset construction
The goal of this module is to conduct a detailed exploration of the structural diversity of perovskite compounds, with the aim of feeding subsequent ML models to map XRD data to materials information. By leveraging extensive crystallographic data repositories, this research seeks to uncover the broad range of structural flexibility and variability inherent to the perovskite family. Perovskites, characterized by the ABX3 stoichiometry, exhibit remarkable structural versatility. Central to their structure is the B site, encapsulated within BX6 octahedra, which can adopt various arrangements and undergo multiple distortions and displacements. The inherent flexibility of the perovskite lattice allows for a wide array of cations and anions to be incorporated, theoretically enabling any combination of atoms to form a potential perovskite structure47.
The initial step is to mine all 93 space groups with ABX3 chemical structure frameworks from the MP database48. These groups extend beyond the typical perovskite space groups conventionally recognized. At the same time, 53 perovskite chemical formulas were found in the ICSD based on real structure.49. This method systematically generates hypothetical perovskite compounds by substituting chemically compatible elements into known perovskite structure templates. A total of 53 representative perovskite compositions extracted from the ICSD database were selected as structural templates. In principle, any composition corresponding to one of the 93 space groups in MP dataset can be mapped to any of these templates for element substitution. The key criterion for element replacement is charge neutrality, following the general formula ABX₃. Since the anion X can be either chalcogen (e.g., O2−, S2−) or halogen (e.g., Cl−, Br−, I−), charge neutrality requires
$$(F)+S(!)+3\times {\rm{valence}}({\rm{X}})=0$$
(1)
To implement this, we developed automated scripts that read CIF files of template structures, identify A- and B-site elements, and replace them according to a predefined list of elements grouped by oxidation state. These substitutions ensure that every resulting structure satisfies Eq. (1). All modified structures were saved as new CIF files, resulting in a virtual dataset of 4929 chemically valid perovskite candidates. The detailed template element replacement List can be found in Table 1.
Data simulation and alignment
This module utilized pymatgen50 to simulate XRD spectra. In the simulation process, element-specific Debye–Waller factors were assigned to account for thermal vibrations and improve spectral accuracy. Wherever possible, these factors were obtained from experimental or computational literature; for elements without reliable references, we substituted values from chemically similar species with the same oxidation state. This strategy ensured internal consistency while mitigating the impact of missing parameters. A complete list of Debye–Waller factors is provided in Table 2. The VSS are essential for expanding the structural hypothesis space and enabling the model to learn robust spectrum-structure mappings. However, a gap exists between real structure spectral data and virtual structure spectral data, which can negatively impact the performance of deep learning models. Therefore, this study aims to help bridge this gap, allowing VSS to more effectively support the analysis of RSS. This strategy was crucial for enhancing the realism of virtual structure spectral datasets, making them more representative of actual conditions.
A fundamental component of this approach was the simulation of unintended phases or contaminants, which are often encountered in experimental samples. By introducing random impurity peaks into the VSS, this study created datasets that were more realistic and challenging, aligning closely with real-world scenarios. This targeted augmentation was vital for robust model training and substantially enhances predictive accuracy on experimental spectra.
In addition to simulating impurity peaks, this study varied the intensities of diffraction peaks to mimic the effect of preferred crystallographic orientation, or ‘texture’. This was done by randomly adjusting peak intensities up to ±50%, to represent the varied orientations of crystallites typically found in real samples. The impact of crystal size on peak broadening, a prevalent phenomenon in XRD analysis, was accounted for by simulating a range of grain sizes employing the Scherrer equation. The simulation spanned from very fine grains resulting in broad peaks to coarser grains producing narrower peaks.
Furthermore, lattice strain-induced peak shifts were introduced by modifying the lattice parameters of each phase, with shifts of up to ±3%. This effectively mimicked the stress and strain effects commonly seen in real samples, which can significantly alter peak positions. Lastly, uniform shifts were applied to all peaks in a spectrum to represent consistent changes in lattice parameters due to factors like temperature variations or chemical composition changes. Crucially, these five methods are not sequential but parallel approaches in this data simulation strategy36.
The resultant VSS, having undergone these augmentation processes, was then combined with RSS to generate synthetic spectra data for model training. To facilitate model evaluation, the RSS were grouped by class labels and stratified sampling was performed to construct multiple subsets with varying proportions. A portion of these subsets was used for data synthesis, while the remaining data were held out in advance for downstream model training and final testing. In this data synthesis algorithm, RSS and VSS are first normalized, respectively5. By randomly selecting one positive sample \({e}_{i}\) from RSS and one negative sample \({t}_{i}\) from VSS, the synthetic sample is created as Eq. (2).
$${X}_{{ac}}={e}_{i}+\varepsilon \cdot {t}_{i}$$
(2)
Where ε is a random number and ε ∈ (0,1).
B-VGG model construction
The model-building process began with the strategic conversion of the VGGNet into a form specifically optimized for processing XRD data. This was achieved by employing a series of 1D convolutional layers with ReLU activation functions, designed to capture the subtle details within the spectra. The model was meticulously optimized to prevent overfitting, with the number of neurons in each convolutional layer being carefully adjusted. This adjustment balanced the extraction of detailed features with the model’s capacity to generalize.
Subsequent max-pooling layers were implemented to reduce dimensionality and distill the most relevant features for classification tasks, further enhancing the model’s efficiency. A key innovation of this model was its structural optimization, as it innovatively integrated Bayesian neural networks (BNNs) to quantify the uncertainty inherent in the predictions. This integration allowed the B-VGGNet to provide not only classifications but also an estimate of prediction uncertainty, which was critical for decision-making in the presence of variability and noise commonly found in XRD data.
The BNN approach introduced a probabilistic framework that assigned weight distributions to the model parameters, capturing both epistemic and aleatory uncertainties. It allowed the posterior distribution of these weights to be estimated using methods such as variational inference, Markov chain monte carlo, or laplace approximation. This feature made the B-VGGNet a robust tool for handling the complexities and uncertainties intrinsic to XRD data analysis.
Uncertainty analysis
Monte carlo dropout (MCD) is recognized as a straightforward yet powerful approach for estimating uncertainty within deep neural networks. The technique hinges on training a model with dropout regularization and maintaining the dropout layers active during the inference phase. By conducting a series of forward passes through the network with varying dropout masks, a collection of predictions is generated, which collectively provide an approximation to the samples from the posterior distribution of the model’s outputs. In this study, VGGNet models were trained with dropout layers strategically placed after each convolutional and fully-connected block. During the inference phase, a multitude of forward passes were executed, each with a unique dropout mask, yielding an ensemble of predictions for each input instance. The aggregation of these predictions, through their mean and variance, provided an estimate of the model’s predictive uncertainty31.
The laplace approximation (LA) is a technique employed for approximating the posterior distribution in Bayesian neural networks, which involves fitting a Gaussian distribution centered on the maximum a posteriori (MAP) estimates of the weights. Initially, the VGGNet models were trained using standard optimization techniques to determine the MAP estimates of the weights. Subsequently, the Hessian matrix of the negative log-posterior concerning the weights was calculated at the MAP estimate. The inverse of this Hessian matrix was utilized as the covariance matrix for the Gaussian approximation to the posterior distribution. This process was repeated with various random initializations and architectures to generate an ensemble of models, culminating in a collection of VGGNet models, each with a LA to its respective posterior distribution51.
Variational inference (VI) is another method used to approximate the posterior distributions that are otherwise intractable in Bayesian neural networks. In this study, VI was applied to approximate the posterior distribution over the weights of the VGGNet models. The goal of VI is to minimize the kullback-leibler (KL) divergence between a computationally feasible variational distribution and the actual posterior distribution. A factorized Gaussian distribution was selected as the variational distribution, characterized by individual means and variances for each weight. These parameters were refined using stochastic gradient descent, to minimize the negative evidence lower bound (ELBO). The ELBO consists of the expected log-likelihood and the KL divergence between the variational and prior distributions. By initiating multiple training sessions with distinct random seeds, an ensemble of VGGNet models was produced, each with an approximated posterior distribution over its weights52.
SHAP value computation for model interpretability
To quantify the contribution of individual diffraction angles to the classification predictions made by the B-VGGNet model, we employed SHAP as a post hoc feature attribution method. SHAP is grounded in cooperative game theory and provides a unified approach to interpret the output of any machine learning model by computing the Shapley value for each input feature.
The Shapley value ϕi for a feature i is defined as the average marginal contribution of that feature across all possible subsets of features. In the context of this study, each input sample is a one-dimensional vector representing the XRD pattern, where each element corresponds to the intensity at a specific 2θ angle. The model output is a probability distribution over the set of space groups, obtained via the softmax function in the final layer of B-VGGNet.
We used the SHAP Python library’s DeepExplainer, designed for deep learning models, to compute these values. The explainer takes the trained B-VGGNet model and a background dataset of reference XRD patterns (typically a subset of training data) as input. For a given sample, SHAP values are computed for each 2θ feature with respect to the predicted class. These values indicate how much the presence of each feature shifts the model output from the expected value (baseline) to the actual prediction.
More formally, for an input sample x ∈ \({{\mathbb{R}}}^{n}\), the SHAP value \({\phi }_{i}\) for the \({\rm{i}}\) feature is computed as Eq. (3).
$${\phi }_{i}=\sum _{S\subseteq F,\{i\}}\frac{|S|!(|F|-|S|-1)!}{|F|!}\left[\,f(S\cup \{i\})-f(S)\right]$$
(3)
Where \(F\) is the set of all features, \(S\) is a subset of features excluding \({\rm{i}}\), and \(f(S)\) denotes the model prediction when only features in subset \(S\) are present.
After computing SHAP values for each sample, we obtained a matrix of shape \(N\times M\), where \(N\) is the number of input samples and\(M\) is the number of 2θ points. These values were used for further statistical aggregation and visualization of feature importance across different space groups.
link








