Helixer: ab initio prediction of primary eukaryotic gene models combining deep learning and a hidden Markov model

Overview of Helixer
Helixer is a deep learning-based framework for eukaryotic gene annotation directly from genomic DNA. It uses a sequence to label neural network that predicts base-wise genomic features including coding regions, untranslated regions (UTRs) and intron–exon boundaries based solely on nucleotide sequence. The architecture integrates convolutional and recurrent layers to capture both local sequence motifs and long-range dependencies, followed by a biologically informed decoding step that assembles coherent gene models. Trained end-to-end on high-quality reference annotations, Helixer generalizes across species without requiring transcriptomic or homology-based evidence. This design enables consistent annotations while minimizing manual curation, providing a scalable solution for annotating newly sequenced genomes and supporting large-scale comparative genomics.
The released Helixer models show state-of-the-art performance compared to existing ab initio gene calling tools and previous Helixer models
As our previous work26 set the state of the art for base-wise predictions, we first compared the latest models to models trained with the hand-selected six (vertebrate) or nine (plant) species used previously, as well as to all intermittently released models (Supplementary Figs. 1–8 and Supplementary Tables 1–4). The best models released here (vertebrate_v0.3_m_0080, invertebrate_v0.3_m_0100, land_plant_v0.3_a_0080 and fungi_v0.3_a_0100), had the highest median Genic F1 for their phylogenetic target range, and showed a more balanced performance across said range, compared to other Helixer models. Nevertheless, no single model was consistently better for all species, so all plotted models are released to allow researchers to select the model likely to perform best for their species of interest.
While promising, the high base-wise performance of raw HelixerBW predictions does not necessarily imply that the performance is maintained through postprocessing. Therefore, we additionally computed the F1 for the phases (phase F1) (Supplementary Figs. 9–13 and Supplementary Table 5), the F1 for the subgenic classes (subgenic F1) (Supplementary Table 6) and the F1 for the genic classes (genic F1) (Supplementary Table 7) scores for the final gene models output by HelixerPost on the selected test species. Importantly, we see that predictions postprocessed via HelixerPost have very similar performance to the HelixerBW predictions, with a slight increase in 43 of 45 test species.
Additionally, we inferred gene models using two existing HMM tools, GeneMark-ES and—where trained models were available for the test species or a closely related species—AUGUSTUS. GeneMark-ES was used without repeat masking, while AUGUSTUS was used both with and without repeat masking.
The phase F1 of HelixerPost (Table 1 and Supplementary Table 5) was notably higher than GeneMark-ES and Augustus across both plants and vertebrates and still somewhat higher for invertebrates generally, although not for all species. All three tools showed similar performance for fungi, with HelixerPost having a slight margin of 0.007 overall. Similar patterns were found with subgenic and genic F1 metrics (Supplementary Figs. 14 and 15 and Supplementary Tables 6 and 7).
Moving from base-wise to feature- or protein-level evaluation, we found lower absolute precision, recall and F1 scores (Fig. 1, Table 2, Extended Data Fig. 1, Supplementary Fig. 16 and Supplementary Tables 8–11). Helixer scored highest for plants and vertebrates, but both HMM tools gained an edge in fungi. Helixer maintained a small advantage in invertebrates overall, although AUGUSTUS and GeneMark-ES were again strongest in some species. Overall, the gene precision and recall scores were lower for every tool than the exon scores, which is expected for the harder task. For most species, Helixer tends to have a higher recall score than precision.
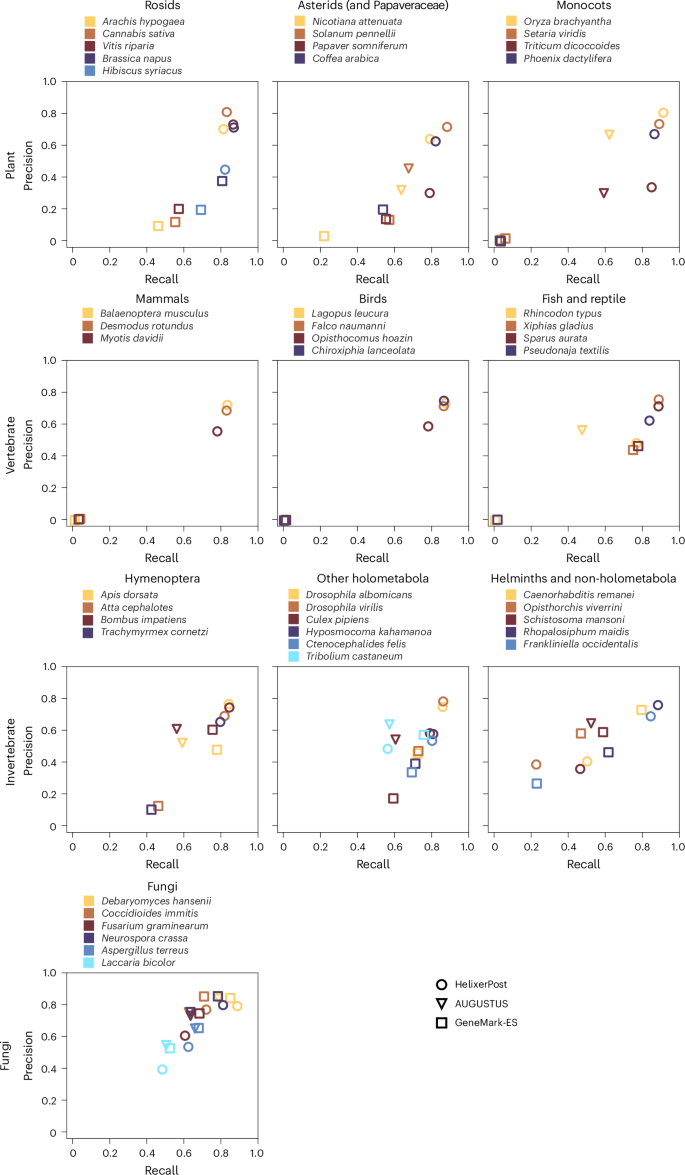
A comparison of precision and recall between HelixerPost (circles), AUGUSTUS (triangles) and GeneMark-ES (squares). The first row shows plants, the second vertebrates, the third invertebrates and the fourth fungi.
When comparing Helixer and AUGUSTUS with softmasking enabled, Helixer outperforms AUGUSTUS in exon F1 score for 9 species, gene F1 score for 10 species and intron F1 score for 8 species out of a total of 16 tested (Supplementary Fig. 17 and Supplementary Tables 8–11).
The completeness of predicted proteomes27 was quantified for the reference annotation and the three prediction tools (Extended Data Table 1, Supplementary Figs. 18–22 and Supplementary Table 12).
The reference annotations generally had the highest performance (39 of 45 species), which is unsurprising since they typically benefit from additional data sources and manual curation. The results of the prediction tools were broadly similar to the base-wise and feature assessments above, with HelixerPost leading strongly within plants and vertebrates, where it even approached the result of the reference. Invertebrates again varied by species, with Helixer leading by a smaller margin overall but GeneMark-ES performing best on several species. Fungi was the most competitive clade, with Helixer leading by a small margin, and interestingly all tools outperformed the reference.
Interestingly, the three invertebrates where HelixerPost scored behind GeneMark-ES in phase F1 and Benchmarking Universal Single-Copy Orthologs (BUSCO) count had the overall lowest BUSCO count in the reference annotations. This hints at larger challenges related to annotating these genomes, be it exceptional divergence or simply a paucity of well-annotated genomes in close phylogenetic proximity to use either for training or for the homology mapping step of an annotation pipeline. However, it may also simply reflect that the invertebrate prediction models are less optimized.
Comparison with Tiberius
To compare Helixer to Tiberius, we trained a model focused on the mammalian clade. We used the three test species Homo sapiens, Bos taurus and Delphinapterus leucas. Even though we could improve Helixers prediction quality compared to our vertebrate model, Tiberius still outperforms Helixer (Fig. 2 and Supplementary Table 13). Especially gene recall and precision is consistently 20% higher. Regarding exon recall, Helixer and Tiberius are almost on par, with Tiberius still taking the lead. For exon precision Helixer shows 10–15% lower values. While we acknowledge that Tiberius outperforms Helixer in the Mammalia clade, we offer a more phylogenetically diverse range of models, especially for the often-underrepresented plant species.
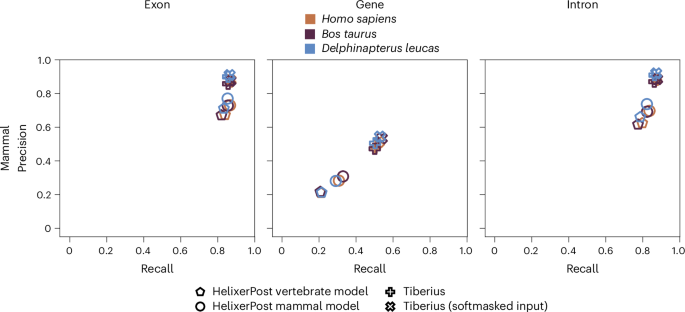
A comparison of precision and recall between the HelixerPost vertebrate model (pentagons), HelixerPost mammal model (circles), Tiberius (pluses) and Tiberius with softmasked input (crosses). The first column depicts exon, the second gene and the third intron metrics.
Ablation analyses
To validate the importance of major changes made during development, we performed an ablation analysis by individually excluding each of the major changes, training a new model, then predicting and comparing prediction quality to the final model (Supplementary Fig. 23).
While none of the changes had a major effect on the overall genic F1 score (Supplementary Fig. 23a), this was not the target of most of the changes. Without transition weights (Supplementary Fig. 23e), the network displays high uncertainty immediately around start and stop codons, as well as acceptor and donor splice sites. All of the low, medium and final transition weights push the networks toward sharper transitions in an example gene, both right at the start codon (Supplementary Fig. 23e) and more widely helping to clear up uncertainty throughout the UTR (Supplementary Fig. 23d), with the final transition weights being most effective. To quantify this on a larger scale, we calculated the genic F1 specifically for the base pairs immediately before or after transitions. Higher transition weights resulted in the expected increase in transition genic F1, with the largest difference between low transition weights and none (Supplementary Fig. 23c). Moreover, the models with reduced transition weights show markedly more internal phase mistakes, that is, where both reference and predictions have a 0–2 codon phase, but the phases do not match (Supplementary Fig. 23b).
Interestingly, including the phase also improved the performance around transitions in the example (Supplementary Fig. 23d,e). In addition, phase predictions are useful signals for postprocessing into final gene models. The long short-term memory (LSTM) model and the hybrid model have similar performances. We acknowledge that this analysis could be made more conclusive by addressing the effect of random starting parameters by adding replicates or, where applicable (the number and arrangement of trainable parameters is only constant for the transition weight ablations), recording and reusing a random seed.
Helixer’s annotations approach reference quality
The F1 metrics shown so far treat the references as the ‘ground truth’; however, the provided references themselves are ultimately the output of a gene annotation pipeline, that is. data-supported predictions. Measuring performance against such references is particularly limited for understanding how good the absolute performance is as it approaches that of the provided reference. In the BUSCO analysis above we already observed that Helixer’s performance was close to that of the reference. Therefore, we used two additional homology-based methods to evaluate in more detail the quality of Helixer and GeneMark-ES predictions versus the reference for the plant test species. AUGUSTUS predictions were omitted since they covered only four plant species.
First, we used the ability of a set of proteomes to form orthogroups as a reference-free indicator for quality, which is particularly relevant when considering annotation applicability for comparative genomic tasks. The more accurate annotation is expected to have more orthogroups with all 12 species represented and generally more orthogroups with higher numbers of species represented. The results are shown in Extended Data Fig. 2 and Supplementary Figs. 24–28.
Here, the reference performs best, with 0.38% of orthogroups containing all 12 species, and 36.5% containing two or more species, followed by Helixer at 0.26% and 31.1% for the same statistics, respectively, and GeneMark-ES at 0.019% and 21.2%, respectively. The reference has the most orthogroups with four or more species, followed by Helixer with two or three species and GeneMark-ES with only one species (Extended Data Fig. 2a). Notably, Helixer’s performance by these metrics was more like that of the reference, that is, to extrinsic data-supported predictions, than to GeneMark-ES’s predictions, which, comparable to Helixer’s are made from the DNA sequence alone. Moreover, all of the reference annotations were respectable (minimum complete BUSCOs of 96.5%, and eight of the species above 99%).
Second, given that the orthogroup-based numbers could potentially be skewed by consistent cross-species mistakes, such as misannotation of transposons, we used the Mapman4 protein annotations (curated plant protein-coding gene families) to evaluate annotation quality. This further gives us a proxy for precision (the percentage of proteins that could be annotated) and recall (the percentage of gene families occupied by a protein). Measured this way, the reference had an average precision, recall and harmonic mean (of precision and recall) of 0.966, 0.931 and 0.948, followed by Helixer (0.878, 0.958 and 0.914, respectively) and GeneMark-ES (0.719, 0.742 and 0.724, respectively) (Extended Data Fig. 2b and Supplementary Table 14).
Thus, the ab initio annotations from Helixer show higher recall than the references for the plant species test set but are behind the references on precision, particularly for the specific species Papaver somniferum and Triticum dicoccoides (Extended Data Fig. 2b, Supplementary Figs. 24, 26 and 28, and Supplementary Table 14), which we note have the most fragmented assemblies, leading to an idea on how to further improve Helixer’s predictions (Supplementary Information).
Filling gaps in Arabidopsis
thaliana
We compared the annotations produced by Helixer for the model plant A. thaliana, to the established high-quality reference annotations TAIR10 and Araport11 (newest versions available in November 2022). When comparing Helixer to Araport11 (Extended Data Fig. 3a), the majority (~70%) of gene loci were found to have an exact match in Helixer. From the remaining ~30%, around 27% are highly similar but varied in their length. The unmatched sequences that remained from Helixer and Araport11 were considered as true annotations if the sequences were found to be conserved in at least 20 streptophyte species. For the Araport11 annotations, 93 of 1,401 were found to be true annotations, while 102 of 711 Helixer annotations were identified as true. Similar results were found when comparing Helixer to TAIR10 annotations (Supplementary Figs. 29 and 30).
From these 102 Helixer annotations not found in the Araport11 reference annotation, a notable example is the Phosphatidylinositol N-acetylglucosaminyltransferase γ subunit. This complex is known to be active in A. thaliana28,29 and the γ subunit locus identified by Helixer shows expression (Extended Data Fig. 3b and Supplementary Fig. 31). However, this γ subunit is entirely missing from TAIR10, and has a chimeric annotation with the next gene in Araport11, resulting in a mere 4 bp of out-of-frame overlap between the resulting protein annotations. While the final postprocessed annotation from Helixer has truncations in at least UTR and introns relative to both the RNA-sequencing data and the raw predictions,the resulting protein was long enough for homology-based identification. This highlights the power of Helixer to complement and improve even the most polished reference annotations.
Benchmarking
As shown in Extended Data Fig. 4, Helixer is faster than the state-of-the-art tools AUGUSTUS and GeneMark-ES, and scales approximately linearly with genome size within phylogenetic groups. When comparing to GeneMark-ES and Augustus on the smallest and largest fungus test genomes and smallest test plant genome, Helixer required 6.2–20.1 fold lower wall time in single-threaded mode. Both AUGUSTUS and Helixer can be parallelized by splitting the genome into multiple fasta files. However, we acknowledge that the other tools make use of more complete multithreading than Helixer does. In single-threaded mode, Helixer.py can annotate the 263-Mbp Oryza brachyantha genome in 27 min and the 3.3-Gbp human genome in just under 8.5 h (Supplementary Methods). The exact speed will vary by system, particularly in regards to the speed of the disk storage and the GPU (Supplementary Fig. 32). A speed comparison to Tiberius was performed by the authors of Tiberius25. In their setup, Tiberius annotated the human genome in 1 h and 39 min and Helixer in 8 h and 54 min.
link








