Flexynesis: A deep learning toolkit for bulk multi-omics data integration for precision oncology and beyond
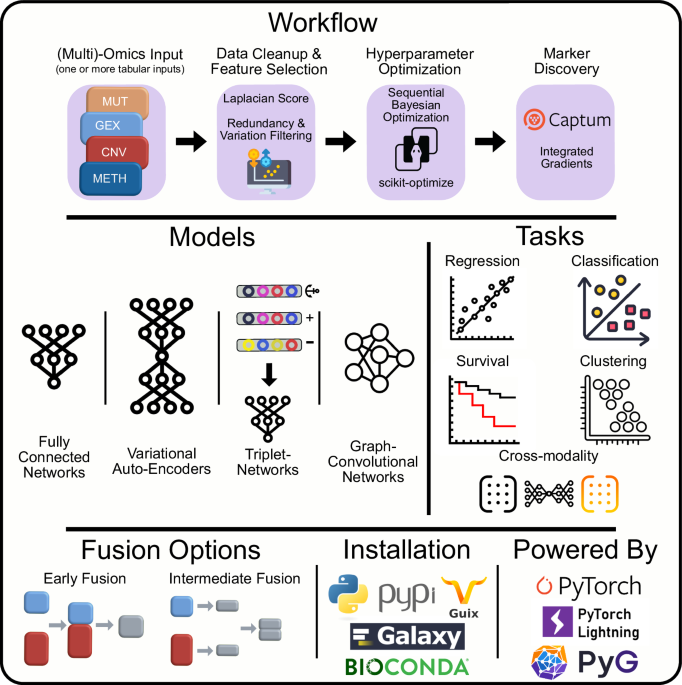
We designed Flexynesis for automated construction of predictive models of one or more outcome variables. For each outcome variable, a supervisor multi-layer-perceptron (MLP) is attached onto the encoder networks (a selection of fully connected or graph-convolutional encoders) to perform the modeling task. Clinically relevant machine learning tasks such as drug response prediction (regression), disease subtype prediction (classification), and survival modeling (right-censored regression) tasks are all possible as individual variables or as a mixture of variables, such that each outcome variable has an impact on the low-dimensional sample embeddings (latent variables) derived from the encoding networks (See Supplementary Figs. 1–8 for the schematic representation of different model architectures, workflows for data processing, hyperparameter optimisation, and model fine-tuning).
Single-task modeling: predicting only one outcome variable
In Fig. 2, we demonstrate the different kinds of modeling tasks that are possible with Flexynesis using a single outcome variable (single MLP) as regression (Fig. 2A), classification (Fig. 2B), and survival models (Fig. 2C). For the regression task, we trained Flexynesis on multi-omics (gene expression and copy-number-variation) data from cell lines from the CCLE database10 to predict the cell line sensitivity levels to the drugs Lapatinib, a tyrosine kinase inhibitor, and Selumetinib, a MEK inhibitor. We evaluated the performance of the trained model on the cell lines from the GDSC2 database18 which were also treated with the same drugs, where we observed a high correlation between the known drug response values and the predicted response values for both drugs (Fig. 2A).

For all three tasks, both a fully-connected-network and a supervised variational auto-encoder was trained and best-performing model’s results were presented. A Performance evaluation of Flexynesis on drug response prediction of a model trained on 1051 cell lines from CCLE (using RNA and CNV profiles) and evaluated on 1075 cell lines from GDSC2 for the drugs Lapatinib (Pearson correlation test, r = 0.6, p = 7.750175e-42) and Selumetinib (Pearson correlation test, r = 0.61, p = 3.873949e-50). The x-axis depicts observed drug response values (AAC-recomputed as in Pharmacogx package)60 and the y-axis depicts the predicted drug response values for the test samples. B Evaluation of Flexynesis on microsatellite instability (MSI) status prediction using gene expression and/or promoter methylation data from seven different TCGA cohorts (gastrointestinal and gynocological cancers) with microsatellite instability (MSI) annotations: TCGA-COAD (Colon Adenocarcinoma), TCGA-ESCA (Esophageal Carcinoma), TCGA-PAAD (Pancreatic Adenocarcinoma), TCGA-READ (Rectum Adenocarcinoma), TCGA-STAD (Stomach Adenocarcinoma), TCGA-UCEC (Uterine Corpus Endometrial Carcinoma), TCGA-UCS (Uterine Carcinosarcoma). The models were trained on 70% of the samples (N = 1133) with MSI status annotations and evaluated on the remaining 30% of the samples (N = 283). The tSNE (t-distributed Stochastic Neighbor Embedding) plot represents the sample embeddings colored by MSI status and the ROC curve represents the best performing deep learning model based on both gene expression and methylation data. C Evaluation of Flexynesis on a survival modeling task on a merged cohort of LGG (Lower Grade Glioma) and GBM (Glioblastoma Multiforme) (using mutations and copy-number-alteration profiles). The model is trained on 557 samples and evaluated on 239 test samples. The tSNE plot depicts the sample embeddings obtained from the model encoder for the test samples colored by the predicted Cox proportional hazard risk scores stratified into “high-risk” and “low-risk” based on the median risk score. The Kaplan-Meier-Plot represents the survival stratification of the test samples based on this risk stratification (Logrank Test, p = 9.94475168880626e − 10). Source data are provided as a Source Data file.
For the single-variable classification task, we demonstrate classification of seven TCGA datasets including pan-gastrointestinal and gynecological cancers with respect to their microsatellite instability (MSI) status using gene expression and promoter methylation profiles. MSI is a molecular phenotype that displays a high mutational load that results from deficient DNA mismatch repair mechanisms19. Moreover, high-MSI levels are predictive of response to immune checkpoint blockade therapies20, underscoring the relevance of detecting MSI-High samples. As MSI-High is characterized by a high mutational load, it would not be surprising to achieve a good classification performance to predict the MSI status using mutation data. We demonstrate that, without using the mutation data, we can achieve a very high accuracy classifier (AUC = 0.981) using gene expression and methylation profiles (Fig. 2B). We have also benchmarked multiple deep learning architectures and data type combinations and observed that the best performing model was trained on gene expression data only (Supplementary Data 2). This result suggests that samples that have been profiled using RNA-seq, but lack genomic sequencing data could still be classified in terms of MSI status.
As the third type of modeling task, we demonstrate survival modeling using Flexynesis on a combined cohort of lower grade glioma (LGG) and glioblastoma multiforme (GBM) patient samples21. For survival modeling, a supervisor MLP with Cox Proportional Hazards loss function is used to guide the network to learn patient-specific risk scores based on the input overall survival endpoints as has been demonstrated previously22. After training the model on 70% of the samples, we predicted the risk scores of the remaining test samples (30%) and split the risk scores by the median risk value in the cohort. The embeddings visualized based on the median risk score stratification shows that the test samples are clearly separable in the sample embedding space, which is also confirmed by the Kaplan-Meier survival plot, which shows a significant separation of patients in terms of predicted risk scores (Fig. 2C).
Multi-task Modeling: Joint prediction of multiple outcome variables
While being able to build deep learning models with any of the regression/classification/survival tasks individually offers an improved user experience, this is also usually possible with classical machine learning methods. The actual flexibility of deep learning is more evident in a multi-task setting where more than one MLPs are attached on top of the sample encoding networks, thus the embedding space can be shaped by multiple clinically relevant variables. This flexibility is even more pronounced in the presence of missing labels for one or more of the variables, which is tolerated by Flexynesis.
To demonstrate the use of multi-task modeling, we trained models on 70% of the METABRIC dataset (a metastatic breast cancer cohort with multi-omics profiles of 1865 patients)23 and obtained the embeddings for the 30% of the samples. In order to compare and contrast the effect of multi-task modeling with single-task modeling, we chose two clinically relevant variables for this cohort: subtype labels (CLAUDIN_SUBTYPE) and chemotherapy treatment status (CHEMOTHERAPY). We built three different models: a single-task model using only the subtype labels (Fig. 3A), a single-task model using only the chemotherapy status of the patients (Fig. 3B), and finally a multi-task model using both subtype labels and chemotherapy status as outcome variables (Fig. 3C). Coloring the samples by the subtype labels and chemotherapy status, we can observe that the sample embeddings obtained exclusively for the subtype modeling reflect a clear clustering of samples by subtype, but not by the chemotherapy status (Fig. 3A). Similarly, the sample embeddings obtained from the model trained exclusively with the chemotherapy status as outcome variable shows a clear separation of samples by treatment status, however the separation by subtypes is not as evident anymore (Fig. 3B). In the multi-task setting where the model had two MLPs (one for subtype labels and one for chemotherapy status), the sample embeddings show a clear separation of both by the subtype labels and also the chemotherapy status (Fig. 3C).

These plots compare the impact of single-task and multi-task modeling on the clustering of samples by clinical variables. Plots on the left are colored by the breast cancer subtype and the plots on the right are colored by the treatment status. A Single-Task Model – Breast Cancer Subtypes: t-SNE (t-distributed Stochastic Neighbor Embedding) visualization of sample embeddings obtained from a single-task model trained exclusively to predict breast cancer subtypes. B Single-Task Model – Chemotherapy Status: t-SNE plot visualization of sample embeddings from a model trained only to predict the chemotherapy status of patients, showing the segregation capability of the single-task model with respect to treatment status. C Multi-Task Model – Subtypes and Chemotherapy Status: t-SNE plot of sample embeddings from a multi-task model trained with dual supervisor heads: one for breast cancer subtypes and another for chemotherapy status. The plot shows how multi-task learning influences the embedding space, enhancing the separation of samples based on both clinical variables simultaneously. Source data are provided as a Source Data file.
We also analyzed the LGG and GBM cohort (from Fig. 2C) in a multi-task setting where we attached three separate MLPs on the encoder layers: a regressor to predict the patient’s age (AGE), a classifier to predict the histological subtype (HISTOLOGICAL DIAGNOSIS), and another survival head to model the survival outcomes of the patients (OS_STATUS). Concurrently training the model with three different tasks at the same time, we inspected the sample embeddings and observed that older patients with high risk scores have the glioblastoma subtype, while younger patients with lower risk scores have the other subtypes, where low risk young patients can still be distinguished mainly by histological subtype (Fig. 4A). Thus, training the model on three clinically relevant variables helps us obtain sample embeddings that reflect all three variables in a hierarchical manner. Inspecting the top markers for each of these variables, we observe common genes for all three variables such as IDH1, IDH2, ATRX, PIK3CA, and EGFR (Fig. 4B), which could be explained by the fact that the clinical variables such as age and histological subtype are correlated with the survival outcomes of the patients, underpinning the importance of these genes in the etiology of the gliomas, which have been extensively studied and reported before24.

The model was trained on 557 training samples from the merged cohort of the LGG (Lower Grade Glioma) and GBM (Glioblastoma Multiforme) patient samples with three supervisor heads: a regressor for the patient age (AGE), a classifier for the histological diagnosis, and a survival head for the overall survival status of the patient (OS_STATUS). A Displays the tSNE (t-distributed Stochastic Neighbor Embedding) visualization of the sample embeddings for 239 test samples, where the size of the points reflect the age of the patient, the colors represent the histological diagnosis, and the samples were stratified into high-risk and low-risk groups based on the predicted risk scores for each patient. The sample embeddings reflect the impact of all three clinical variables concurrently. B Displays the top 10 most important features discovered for each supervisor head for the patient’s age, histological subtype, and survival status. Source data are provided as a Source Data file.
Unsupervised learning: finding groups and general patterns
One of the main architectures provided in Flexynesis is the variational auto-encoders (VAE) with maximum mean discrepancy (MMD) loss25. While VAEs are usually employed in unsupervised training tasks, in Flexynesis they can be used for both supervised and unsupervised tasks. In the absence of any target outcome variables (in other words, without any additional MLP modules attached on top of the encoders), the network behaves as a VAE-MMD where the sole goal is to reconstruct the input data matrices, while generating embeddings that follow a Gaussian distribution due to the MMD loss.
As a proof of principle experiment, we trained a VAE-MMD model without any attached supervisor MLPs, to test the unsupervised dimension reduction capabilities on 21 cancer types from the TCGA resource using gene expression and methylation as input modalities. Applying k-means clustering (k from 18 to 24), we obtained a clustering of the samples based on the trained sample embeddings. The tSNE representation of the resulting sample embeddings shows a clear separation of unsupervised clusters (Fig. 5A) and the known sample labels (Fig. 5B) with a good correspondence between unsupervised clusters and known sample labels (adjusted mutual information: 0.78) (Fig. 5C, Supplementary Data 3).

The figure displays the unsupervised analysis of 21 cancer types from the TCGA study for 1600 samples (80 samples per cancer type were randomly selected). A Displays the tSNE plot of the training sample embeddings colored by the best performing clustering scheme using the k-means algorithm for values of 18 < = k < = 24, where best clustering was selected by the best silhouette score (k = 24). B The same tSNE (t-distributed Stochastic Neighbor Embedding) plot as in (A) but colored by the known cancer type labels. C The heatmap displays the concordance between the cluster labels from (A) and known cancer type labels from (B), where the adjusted mutual information score is 0.78. Each row is normalized to add up to 100% where the color of the cells represent the concordance percentage of the cancer types to the corresponding cluster labels. Source data are provided as a Source Data file.
Cross-modality learning: transferring knowledge between different omic data types
While variational autoencoders are designed to reconstruct the initial input data, this can be formulated in a different fashion such that the goal of the reconstruction is a set of matrices different from the inputs. Thus, it is possible to build models where the input data modalities differ from the output data modalities. For instance, a gene expression data matrix could be used to reconstruct a mutation data matrix, thus learning how to translate between these modalities, while simultaneously learning the low-dimensional embeddings that reflect this translation. Due to the modular structure of the Flexynesis, we can also attach one or more MLPs on top of these cross-modality encoder models for one or more target variables as supervisors for regression, classification, and survival tasks.
In order to demonstrate this feature, we designed an experiment using the genome-wide gene essentiality scores measured for >1000 cell lines as part of the DepMap project26. The DepMap database contains measurements of cellular proliferation after perturbation of all protein coding genes. It has been previously shown that for a given cell line, the gene expression profiles of the cell lines can be used to predict the gene essentiality scores27,28. Here, we carried out a similar approach, where gene expression profiles of genes across cell lines were used as input with a goal to reconstruct the cancer cell line dependency scores of the same genes. We expanded this approach to a multi-modal setup, where we used two additional data modalities besides the gene expression: (1) we used pre-trained large language models to generate protein sequence embeddings for the same genes using Prot-Trans29 and obtained sequence embedding vectors for each gene (using the canonical protein sequences) (2) we used the structural and functional features of proteins (such as disorder profiles, evolutionary sequence conservation, secondary structures, post-translational modification sites) from the DescribePROT database30. Thus, each gene was represented by three data modalities: gene expression profiles across cell lines, protein sequence embeddings, and describeProt features. We used these modalities to reconstruct the gene-essentiality scores for each of the cell lines in the DepMap database. In addition, we attached a supervisor MLP to guide the network to predict the hubness-score of each gene in the genetic interaction networks obtained from the STRING database31 assuming that the centrality of a gene in biological interaction networks could be a contributing factor in its essentiality for cell survival (Fig. 6A). Thus, the model was trained concurrently to predict both the gene essentiality score in a particular cell line (as a matrix), along with the gene hubness (as a vector). We trained the model on 70% of the genes and evaluated the model on the remaining 30% of the genes, by computing the average correlation of each cell line’s predicted gene dependency scores with the measured scores. The addition of the protein sequence embeddings from the language models had a significant improvement on the performance of the model, while the addition of DescribeProt features did not make an additional improvement over the protein language embeddings (Fig. 6B), which suggests that LLM-based protein embeddings might be already capturing similar information to the features from describePROT. For comparison, we also built models without the supervision for the “hubness” feature. We observed that depending on the data combinations used as input, using a supervisor for “hubness” led to an improvement for the single data modality case (Fig. 6B, right panel) but also a deterioration for the reconstruction scores when all three modalities were used as input (Fig. 6B, middle panel), which could be because the network may have put more weight on learning the “hubness” feature while the weights on cross-modality reconstruction may have been diluted with the addition of further data modalities in this particular case.

A Multi-modal cross-modality prediction of gene knock-out dependency probabilities of cell lines. A cross-modal encoder-decoder model was used that takes as input a combination of input data modalities and reconstructs the CRISPR-based gene knock-out dependency probability scores of cell lines from the DepMap project. Each gene is represented with a combination of feature sets including the expression profile of the gene across cancer cell lines, Prot-Trans large language model embeddings of the gene’s canonical protein sequence, and functional/structural sequence features of the gene’s canonical protein sequence from the describePROT database. The cross-modality encoder was trained both with and without an attached supervisor that predicts the hubness score of the gene according to its centrality in the STRING database. B The distribution of the correlation scores for each (N = 1064) cell line’s measured gene knock-out dependency scores and the predicted scores (for the test set) based on different input data modality combinations: “Gene Expression” represents the prediction performance when using only gene expression profiles of the genes in the cell lines; “Gene Expression + Protein embeddings” represents the prediction performance when using both the gene expression profiles and protein sequence embeddings from Prot-Trans; “Gene Expression + Protein embeddings + DescribeProt Features” represents the prediction performance when using all three feature sets including gene expression, protein embeddings, and describeProt features. The center line in each violin represents the median, and the shaded regions span the interquartile range (IQR). Individual data points each representing one cell line (N = 1064) are overlaid using jittered dots, color-coded by hubness category. Source data are provided as a Source Data file.
Improving model performance via model fine-tuning
One of the conveniences offered by neural networks compared to classical machine learning approaches is that the neural networks trained on a source dataset can be fine-tuned on a small portion of the target dataset. This feature offers a possibility to tune the trained model on the potentially shifted distribution of the target dataset compared to the source32. We implemented an optional fine-tuning procedure, which uses a portion of the test dataset to modify the model parameters (following a combination of model parameter freezing strategies and different learning rates). The fine-tuned model is then evaluated on the remaining test dataset samples. In the first experiment, we trained multiple neural network models along with baseline methods (Random Forests, SVM, XGBoost) on drug response profiles of the CCLE database and fine-tuned the trained neural network models on 100 samples from the test dataset (GDSC database). We observed that, while fine-tuning can be beneficial for different models, it doesn’t create an overall meaningful difference from the models that were not fine-tuned (Fig. 7A, see Supplementary Data 4 – Sheet 2 for paired bootstrap test statistics).

Violin plots show the distribution of performance values estimated from 100 bootstrap replicates. The solid black point indicates the mean, and the error bars represent the 95% confidence interval (CI) computed from the empirical bootstrap distribution. Individual bootstrap values are overlaid as jittered dots, color-coded by method or experimental condition. A Deep learning and classical machine learning methods were trained on the CCLE dataset for Selumetinib response levels (using gene expression and copy number variation data) and evaluated on the GDSC2 dataset. Deep learning models were further fine-tuned using 100 samples from GDSC2 and the models were evaluated on the remaining unseen test samples from the GDSC2 dataset. The best performing fine-tuned deep learning model is compared to other non-fine-tuned deep learning and baseline models. See also Supplementary Data 4 for paired two-sided bootstrap t-test statistics for additional drug response model performance comparisons with/without fine-tuning. B Models were trained on TCGA tumor samples from three different cancer types (breast cancer, glioblastoma, and colorectal cancer) and evaluated on CCLE cancer cells lines derived from the same three cancer types with and without fine-tuning the models using 50 samples during fine-tuning stage. The best performing fine-tuned deep learning model is compared to other non-fine-tuned deep learning and baseline models. See also Supplementary Data 4 for paired two-sided bootstrap t-test statistics. Source data are provided as a Source Data file.
As the CCLE and GDSC databases have a relatively similar origin, resulting in good concordance with similar distributions, fine-tuning didn’t yield a clear advantage. Therefore, we tested fine-tuning in a separate experiment where the source (training) dataset and target (test) datasets come from completely different sources. We built models to predict the cancer types of human tumor samples from three different TCGA cohorts (breast cancer, glioblastoma, and colorectal cancer) and used the trained model to predict the cancer types of cell lines derived from the corresponding three different cancer cell lines from the CCLE using gene expression and copy number variation data as input. We observed that all the models performed very poorly without fine-tuning, with an F1 score of ~0.16, with similar performances by Random Forest, XGBoost, and SVM models, too. However, fine-tuning the deep learning models using 50 samples led to a significant improvement in the prediction performance achieving F1 scores of up to 0.8 (Fig. 7B, see Supplementary Data 4 – Sheet 1 for paired bootstrap test statistics).
Discovering biomarkers of drug response in cell lines
All model architectures implemented in Flexynesis, are equipped with a marker discovery module based on Integrated Gradients and GradientSHAP feature attribution methods33,34,35. In order to evaluate whether the trained Flexynesis models can capture known/expected markers, we constructed models predicting drug response, for eight drugs with known molecular targets. The models were trained on drug response data from CCLE and evaluated on the corresponding features from the GDSC dataset. We trained both a fully connected network (DirectPred), a supervised variational auto-encoder (supervised-vae), and graph-convolutional neural network (GNN-SAGE) using various data type combinations (mutations, mutations + RNA expression, and mutations + RNA expression + copy number variants). The top ten markers per drug were extracted from the best performing model among all the experiments (Fig. 8A) using both Integrated Gradients and GradientSHAP methods. The two methods yielded almost identical results (Supplementary Fig. 9), therefore we report the feature attribution metrics only from the Integrated Gradients method. We labeled the top markers by data type and also by the presence of the marker in civicDB36, a database of clinically actionable genetic biomarkers of drug response. For 6 out of 8 drugs, we could find at least one known marker, present in civicDB (Fig. 8B). In addition, we observe that the best performing models are never trained on “mutations-only”. Top markers for each of the drugs are dominated by single nucleotide variants, however, we also observe that the best performing models (Fig. 8A) are the ones where the mutation data is complemented with at least the “RNA” layer, which is in line with previous findings, we and others have demonstrated before, that using the gene expression data on top of the mutation features significantly improves drug response prediction performance37,38.

A fully connected network (DirectPred), a supervised variational autoencoder (supervised_vae), and a graph-convolutional network (GNN-SAGE) was trained on three combinations of data modalities: using only mutations; using mutations and RNA expression; using mutations, RNA expression, and copy number alterations. A Best performing model+data type combination for each drug is displayed. Color scale (shades of blue to red) reflects the pearson correlation score. B The top 10 markers (in the y-axis) discovered for each drug (based on the best performing model + data type combination depicted in panel (A). The markers are both labeled and colored by the corresponding data modality (dark blue: RNA expression, light blue: Mutation). The markers that are already known to be indicator markers for the corresponding drug according to the CIViC (Clinical Interpretation of Variants in Cancer) database are labeled as “Known Target (CIVIC)”. The x-axis displays the relative importance of the top markers, where the best marker has a value of 1. While most drugs have dominantly mutation markers in the top 10, the best performing models always have RNA expression as an additional data modality. Source data are provided as a Source Data file.
The Flexynesis benchmarking pipeline
Previous benchmarking of different neural network architectures13,14,39, showed that none of the methods outperform others in all tested scenarios. It is challenging to choose the best performing neural network architecture along with the type of multi-omic modalities best suited for a given task ahead of time. Additionally, it is possible that the accuracy of the classical machine learning methods, such as a random forest classifier, is sufficient for a given prediction task. Therefore, to attain the best performing model, we have to execute multiple experiments with different data type combinations, different fusion approaches, and different neural network architectures. Moreover, some tasks might benefit from building multi-task training, while others might perform better for the target variable of interest in a single-task setting.
To accommodate such combinatorial experimentations, we setup a benchmarking pipeline which can be configured to run different flavors of Flexynesis on different combinations of data modalities, different fusion options, fine-tuning options, along with a baseline performance evaluation using random forest, support vector machines, XGBoost and random survival forest methods. The pipeline then builds a dashboard with rankings of different experiments in terms of prediction performances for different tasks.
We ran the benchmarking pipeline on datasets with clinically relevant outcome variables and built a dashboard of rankings of the different experiments (See dashboard and Supplementary Data 6). We designed 14 different tasks across 5 different datasets in a total of 222 different experiments, where we tested different tools, tool flavors, data fusion and fine-tuning options. Immediate observation confirms previous findings that no single neural network model outperforms others in all tasks. Of the 14 tasks, the top ranking method was equally divided between deep learning models and classical machine learning models (SVM, Random Forest, XGBoost) (Fig. 9A, see Supplementary Data 6: Sheet 2 for paired bootstrap tests for the comparison of best performing deep learning and baseline models). Cross-experiment comparison of model performances suggest a slight edge for deep learning models (Fig. 9B) with small effect sizes, nonetheless. Furthermore, we compared the deep learning models in terms of omics data modality fusion options (see Methods: Data modality fusion options). Among the best performing models, we don’t observe a significant difference between early or intermediate fusion settings (Fig. 9C). Similarly, fine tuned deep learning models don’t show any insignificant improvement over the counterparts with no fine-tuning (Fig. 9D). Finally, among the GNN models, the choice of SAGE convolution method yields slightly better results in our experiments (Fig. 9E), but doesn’t achieve statistical significance. These experiments suggest that the choice of deep learning versus baseline methods, or deep learning method settings such as model tuning, fusion options, or convolution methods probably depends on the specific task and it is not possible to generalize to all possible situations. The value of different approaches is task specific, therefore we advise running multiple experiments to obtain the best model for the dataset at hand. This accessory pipeline ameliorates the execution of such experiments. The pipeline is available at https://github.com/BIMSBbioinfo/flexynesis-benchmarks.

The “score” in the y-axis for panels B-E are scaled to 100 (where the best model gets a score of 100) in order to enable comparison of different scoring metrics. Description of box-plots: the center line indicates the median, the box bounds represent the 25th and 75th percentiles, and the whiskers extend to the most extreme values within 1.5 × the interquartile range (IQR). Individual data points are overlaid using jitter to visualize sample density and spread. Statistical comparisons between groups were performed using a two-sided Wilcoxon test. A Best performing model setup for each of 14 different tasks. B Comparison of the best result of among any deep learning method (N = 14) with the best result of the classical machine learning approaches (N = 14) (two-sided Wilcoxon test, p = 0.037) C Comparison of the best performing data fusion strategies: intermediate fusion (N = 12) and early fusion (N = 12) for the deep learning models in experiments where there were at least 2 data modalities (two-sided Wilcoxon test, p = 0.77). D Comparison of the best performing model with (N = 11) and without (N = 14) the fine-tuning option (two-sided Wilcoxon test, p = 0.2). E Comparison of the best performing GNN models in terms of different convolution options: SAGE (N = 14) vs GC (N = 14) (two-sided Wilcoxon test, p = 0.73) and GCN (N = 14) vs GC (N = 14) (two-sided Wilcoxon test, p = 0.55). Source data are provided as a Source Data file. See Supplementary Data 6 for full table of results and bootstrap testing statistics for comparisons of best performing deep learning and baseline models.
link








