Enhanced MRI brain tumor detection using deep learning in conjunction with explainable AI SHAP based diverse and multi feature analysis
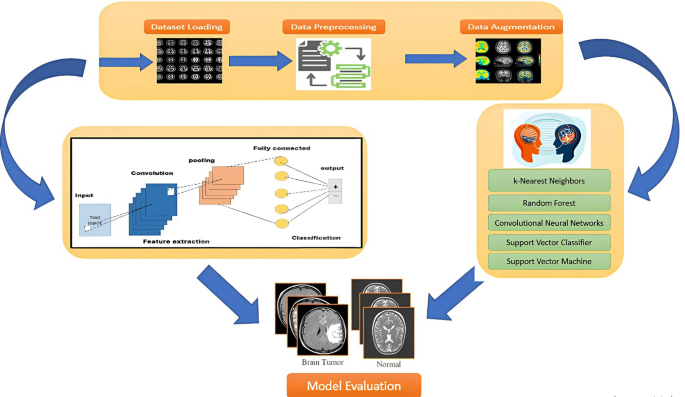
This section discusses comprehensively the performance assessment of various feature representation techniques (CNN, DWT, FFT, Gabor, GLRLM, and LBP) in conjunction with several machine learning and deep learning algorithms. Performance Analysis of Feature Extraction Techniques with ML Classifiers. Class weighting is implemented during training to rectify the class imbalance in the sizable dataset, giving the minority (non-tumor) class larger loss penalties. Furthermore, we reported additional performance metrics that better reflect performance on imbalanced datasets, such as precision and F1-score, in order to ensure a fair evaluation.
CNN feature space
CNN serves as both a feature extraction and classification technique in this work. To extract deep features from the MRI images, we used a pre-trained CNN model (such as imageNet in the manuscript) and frozen the convolutional layers during inference. For the final classification, these features were subsequently fed into different machine learning classifiers (such SVM, KNN, or PNN). The CNN was not fine-tuned or trained end-to-end with the classifier. This fixed-feature extraction approach was chosen to reduce training time and computational cost, and to evaluate the effectiveness of transfer learning in combination with classical classifiers.
In case of classification, a CNN is employed in an end-to-end fashion, where the network simultaneously learns feature representations and performs classification. The convolutional layers act as learnable feature extractors, and the resulting features are passed through fully connected layers to generate the final classification. All layers are trained jointly from scratch without using pre-trained weights.
The CNN-based image representation approach demonstrated remarkable performance across all learning algorithms are reported in Table 2. Among these algorithms, the SVC algorithm outperformed compared to other algorithms, and achieved 96.6% accuracy and 98% specificity, highlighting its effectiveness in identifying MRI-based tumors. So the success rates of SVC are balanced in the case of all used metrics. On the other hand, RF and CNN exhibited high precision and specificity ensuring reliable positive predictions. In contrast, the sensitivity of RF and CNN is unbalanced. So the models are biased toward one class. kNN obtained 81% accuracy and 80% F1-score, while sustaining success rates. While PNN performed moderately achieved 83% accuracy and 88% specificity. In this study, robust features such as spatial hierarchies in images are captured due to CNN capabilities and eradicating the requirements of manual feature engineering. In contrast, it is computationally expensive owing to its complex structure and also shows that it is not optimal for distance-based learning. In CNN the learned attributes are not interpretable, so it is sometimes difficult to make a decision. It is overfit in the case of a small dataset.
Discrete wavelet transform (DWT) feature space
Discrete Wavelet Transform (DWT) yielded robust outcomes in which SVC still led with 95.9% accuracy and 97.8%% specificity ensuring reliable predictions are shown in Table 3. In a sequel, CNN and RF also exhibited strong classification power with 100% precision. The kNN model showed excellent sensitivity of 93% and a balanced F1-score, emphasizing its capability to recognize true positives effectively. In addition, PNN also performed well with 99% sensitivity and 93% accuracy. The significance of DWT is the multi-resolution analysis which extracts both frequency and spatial information. In contrast, it requires careful consideration in the case of wavelet functions and decomposition level selection.
FFT feature space
Fast Fourier Transform (FFT) features obtained optimal success rates using SVC, which are 97.4% specificity and over 95.2% accuracy reported in Table 4. CNN maintained a high precision rate of 93%, while kNN demonstrated effectiveness in true positive detection with is sensitivity of 97%. The sensitivity of RF is high but low specificity which reveals imbalanced classification results. FFT is computationally efficient compared to DWT along with extracting global frequency information. In contrast, it lacks spatial information, which restrains its effectiveness for localized information. FFT is sensitive to noise and assumes the signal is periodic, which may not be always true in real-time applications.
Filters Gabor feature space
The Gabor representation method demonstrated moderated results which are shown in Table 5. In this method, SVC continuously reflected exceptional performance with 93.6% accuracy and 95.4% specificity. CNN provided a commendable precision rate of 90%, while kNN and PNN achieved comparatively lesser accuracy at 74%, indicating that Gabor features may lack of discriminative power of alternative methods. The importance of the Gabor method is the extraction of texture and edge information. On the other hand, it is computationally expensive owing to the multi-scale and multi-orientation nature of Gabor filters. Gabor filter focuses on local information while ignoring global patterns in data.
GLRLM feature space
GLRLM-based information exhibited a robust correlation with SVC, obtaining an accuracy of 94.8% and 96.6% specificity are shown in Table 6. kNN yielded a balance between recall and precision, while CNN and RF successfully delivered correct positive predictions. In addition, PNN obtained 81% of accuracy which performed moderately. The performance variations indicate that GLRLM is appropriate for specific learning algorithms but not universally optimal. However, GLRLM representation methods extract texture information efficiently while its ability is limited to capture spatial relationships in images.
LBP feature extraction
Local Binary Pattern (LBP) features show excellent outcomes which are shown in Table 7. SVC again obtained the highest success rate which is 98.06. The sensitivity and specificity of SVC are balanced. CNN yielded the second-highest accuracy of 97.8%, succeeded by RF at 95%. The balanced sensitivity, precision, and F1-score across learning algorithms validated the robustness of LBP features in brain tumor classification. LBP is effective in collecting local texture patterns and is also efficient and simple computationally.
In order to investigate the contribution of each feature representation scheme, statistical test chi-square and p-value are computed which is illustrated in Table 8. The results indicate that CNN, DWT, and LBP exhibit a significant impact (p < 0.05), thereby confirming their importance in the classification process. The results indicate that FFT, GRLM, and Gabor do not achieve statistical significance (p > 0.05), suggesting a minimal contribution to classification accuracy.
Performance metrics of learning algorithms using large training dataset
The empirical results reveal that leaning algorithms have achieved the highest success rates on LBP feature representation methods. In order to further investigate and highlight the strength of LBP feature representation methods, the large dataset is expressed by the LBP method. In this dataset, the total number of MRI images are 7023 of which 5723 images are for training and 1311 images are for testing. The performance of learning algorithms on LBP feature space is shown in Table 9. The highest accuracy is achieved by CNN with 98.9% while SVC obtained 96.7% accuracy. The sensitivity and specificity of CNN are balanced. On the other hand, the performance of kNN, RF, and PNN are comparatively the same. The performance of learning algorithms on LBP feature space is still advanced compared to other learning algorithms in the case of large training datasets.
The AI explainable SHAP analysis (Fig. 2) revealed that the most influential LBP features correspond to regions with high texture contrast and edge definitions. These align with known MRI characteristics of brain tumors. For instance, uniform texture patterns with high SHAP scores were often associated with meningioma, while features indicative of texture heterogeneity were linked to gliomas. This correspondence affirms the model’s clinical interpretability and the usefulness of the LBP-based features in capturing tumor characteristics that have medical significance. In Fig. 2 each row corresponds to a specific feature, where red points denote higher feature values and blue points indicate lower values. The SHAP values represented on the x-axis quantify the influence of features on model predictions. Positive SHAP values derive predictions toward the tumor, while negative values indicate healthy. Various feature subsets with different dimensions were evaluated, and the optimal selection of the highest-ranked features markedly improved the model’s prediction performance.
In addition, Accuracy of individual classes of tumors on both datasets of CNN and SVC are listed in Table 10.
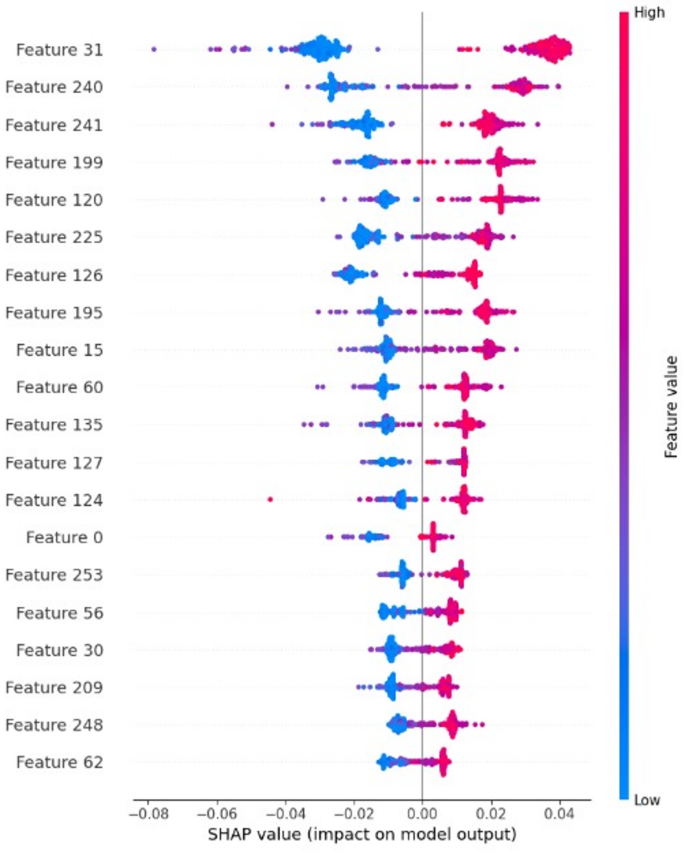
SHAP interpretation of LBP features.
In this work, comparison analysis was also carried out, focused on interpretability, deployment feasibility, and computing efficiency in order to substantiate the advantages of our proposed deep learning model over the transformer-based model. Our proposed model performs excellently compared to the Swin Transformer baseline in terms of computational cost, exhibiting about twice the inference performance on an NVIDIA RTX 3080 GPU, about 65% fewer floating-point operations (FLOPs), and 50% less memory utilization. These enhancements make our proposed model a strong contender for deployment in resource-constrained environments, such as rural healthcare settings where restricted hardware capacity and power availability often prohibit the implementation of more intricate transformer architectures. Additionally, our proposed model CNN-SHAP approach, uses feature attribution maps to produce interpretable visualizations in order to recognize the interpretability and generalizability trade-offs. In medical deep learning, model transparency must guarantee diagnostic reliability without sacrificing clinical acceptability. This conflict has been highlighted in published research by emphasizing the necessity of explainability in high-stakes applications Doshi-Velez and Kim (2017) and Tjoa and Guan (2020). In order to make our method both efficient and useful for actual clinical applications, we strive for a balance between obtaining high performance on benchmark datasets and preserving transparency through SHAP-based post hoc explanations. The performance of the proposed model is evaluated using a stratified five-fold cross-validation test. The mean and standard deviation (mean ± SD) of each class are reported in Table 11. It demonstrates the consistency and reliability of the proposed model on both small and large datasets. In addition, 95% confidence intervals are also computed applying a Student’s t-distribution (df = 4 and sample size n = 5) for each class and shown in Table 12. The narrow confidence intervals indicate high model stability and consistent performance.
Discussion
A comprehensive performance evaluation indicated significant differences in the performance of various combinations of feature extraction methods and classifiers. The SVC exhibited consistently high accuracy and specificity, establishing it as a dependable model for the detection of brain tumors in MRI scans. This indicates that SVC demonstrates a high proficiency in differentiating between tumor and non-tumor cases while maintaining a low rate of false positives. The RF classifier demonstrated high precision, reflecting its effectiveness in producing accurate positive predictions and reducing the occurrence of misclassifications. The CNN model demonstrated significant performance improvements, particularly in minimizing false positive instances (Relu, softmax, Adam), thereby increasing diagnostic reliability. In the meantime, the kNN algorithm demonstrated high sensitivity, successfully identifying true positive cases. This emphasizes its ability to accurately identify the presence of tumors; however, additional optimization may be necessary to minimize false positives.
LBP space has shown significant effectiveness in capturing texture-based features, leading to high classification accuracy. This highlights the significance of selecting a suitable feature representation method to improve tumor detection. Additionally, this study emphasizes the critical importance of feature extraction methods in enhancing diagnostic accuracy within medical imaging applications.
In the case of training dataset1, the prediction outcomes of SVC are reported the highest using all feature spaces. SVC obtained 98.06% accuracy on LBP feature space. The second highest success rates are obtained by CNN and are comparatively the same as SVC. CNN yielded 97.7% accuracy on LBP feature space. Further, evaluating the generalization and resilience of learning algorithms on the LBP feature representation method another large dataset training dataset2 is used. In the case of a large dataset expressed by the LBP feature representation method, CNN was identified as the highest-performing model, with a weighted accuracy of 98.9%, and SVC obtained 96.7% accuracy. In the case of the small dataset, the machine learning algorithm SVC is well performed and the deep learning model CNN is very close in terms of accuracy. In contrast, the predictive outcomes of CNN in the case of large datasets are much better compared to SVC. The empirical results elaborate that as the number of images increased the performance of SVC reduced while the performance of CNN enhanced. So, machine learning algorithms are useful in the case of small datasets, in contrast, CNN performs well in the case of large datasets.
In conclusion, the results highlight the essential relationship between feature extraction methods and machine learning and deep learning classifiers in enhancing the accuracy of brain tumor detection. The findings demonstrate that LBP-based feature extraction yields strong feature representations. However, traditional methods such as DWT and Gabor filters also show competitive performance when utilized alongside effective classifiers like CNN, SVC, and RF. The insights presented have significant implications for enhancing the accuracy and reliability of automated diagnostic systems within the field of medical imaging.
CNN identifies spatial hierarchies of features, effectively capturing patterns including edges, textures, and intricate structures, thereby demonstrating high suitability for image-based tasks. The deep learning architecture enables the achievement of higher accuracy in large and complex datasets. The model demonstrates strong generalization capabilities across diverse image datasets through the acquisition of robust feature representations. The system effectively manages variations including rotation, scale, and intensity differences by utilizing convolutional layers and pooling mechanisms. SVC is dependent on pre-extracted features, indicating that its performance is significantly influenced by the quality and relevance of these features. The system does not acquire spatial hierarchies; instead, it identifies optimal decision boundaries for classification based on the input features. It demonstrates robust performance when supplied with well-extracted and relevant features. It may encounter difficulties when processing high-dimensional and raw image data unless extensive preprocessing is applied. The system exhibits increased sensitivity to variations in image data due to its lack of inherent spatial feature learning, resulting in reduced robustness when addressing diverse and complex image patterns. It is computationally less expensive in comparison to deep learning models. The method demonstrates effective performance on smaller datasets and exhibits ease of training; however, it may not achieve the same level of proficiency as CNN in processing highly complex image data. However, the dependence on extracted features indicates that performance may be suboptimal when encountering unseen or complex variations in medical images.
SVC demonstrates high effectiveness in structured classification tasks that utilize well-engineered features. However, CNN exhibits superior performance in medical image analysis, attributed to its capability to learn intricate spatial features, achieve robust generalization, and uphold elevated accuracy and specificity. Considering the intricacies involved in MRI brain tumor detection, CNN is identified as a more effective option for automated and high-precision medical imaging applications.
Comparative analysis of exiting models with propose model
This study distinguishes itself by employing multiple machine learning and deep learning algorithms in conjunction with diverse feature representation strategies, validating a comprehensive comparative analysis with existing research are reported in Table 13. The empirical results highlight the significance of our proposed approach in enhancing classification performance. For instance, a prior study75 reported an accuracy of 93.3% using a CNN-based model. Similarly, an auto-encoder network and 2D CNN were proposed in76, attaining 95.63% and 96.47% accuracy, respectively. The kNN classifier in77 achieved an accuracy of 86%. A novel hybrid approach combining CNN with classical architectures was proposed in78, yielding 96% accuracy along with 95% sensitivity and precision. Gradient Boosting, identified as the best-performing classifier in79, achieved an accuracy of 92.4%, an F1-score of 89.5%, 94.4% sensitivity, and 85% precision. Another brain tumor classification study80 attained 97.15% accuracy with 97% precision, recall, and F1-score. To further validate the robustness of the model, a new model was introduced BRATS 2018 obtained 94.6%, accuracy, respectively81. In contrast, our proposed model achieved 98.9% accuracy, 98.6% sensitivity, 99.0% precision, and 99.2% specificity.
The proposed model exhibits enhanced performance across all evaluated metrics compared to existing models. The integration of LBP feature space along with CNN for classification likely contributes to its high accuracy, sensitivity, precision, F1-score, and specificity. The improved performance of the model highlights the effectiveness of merging texture-based feature extraction with the deep learning algorithm CNN for MRI-based brain tumor classification. Consequently, it presents itself as a viable option for brain tumor classification tasks, especially in medical applications where minimizing false positives and false negatives is critical. However, further, the proposed model is validated on larger and more diverse datasets in order to confirm its generalizability and robustness.
link








