Domain-incremental white blood cell classification with privacy-aware continual learning

Datasets
We utilize a series of WBC datasets demonstrating challenging domain shifts commonly found in clinical settings, which we refer to as ‘heterogeneous shifts’. These shifts can be attributed to differences in acquisition centers, staining protocols, imaging conditions, and organs (blood smears and bone marrow). We consider four datasets, namely PBC, LMU, MLL, and UKA, which exhibit notable variations.
PBC: This dataset consists of 17,092 images of White Blood Cells (WBCs), which are categorized into 11 distinct classes. This dataset is derived from normal peripheral blood cells8. The smears were prepared with May Grünwald-Giemsa staining using the slide maker-stainer Sysmex SP1000i. The cell images were digitized with CellaVision DM96 to JPG images with size 360 \(\times\) 363 pixels.
LMU: As presented in9, this dataset contains over 18,000 images of single WBCs from peripheral blood smears that have been classified into 5 major types9. The single-cell images were produced by M8 digital microscope/scanner (Precipoint GmbH, Freising/Germany) at 100\(\times\) magnification and oil immersion. Here, the physical size of a camera pixel is 14.14 pixels per \(\mu\)m.
MLL: This dataset is a subset from another publicly available source10. It consists of bone marrow smears from cohorts over 170,000 images of WBCs, encompassing 8 different cell types stained at the Munich Leukemia Laboratory. The bone marrow smears were scanned with a magnification of 40\(\times\), stained with May-Grünwald-Giemsa/Pappenheim staining. The samples were digitized with a Zeiss Axio Imager Z2 into 2452 \(\times\) 2056 pixels, and the physical size of a camera pixel is 3.45 \(\times\) 3.45 \(\mu\)m.
UKA: This is an internal WBC dataset derived from bone marrow smears, comprising 11,899 labeled images of WBCs across 36 distinct classes, collected from 6 patients at Uniklinikum Aachen. The samples were acquired using a Leica Biosystems Aperio VERSA 200 whole-slide scanner with 63\(\times\) magnification and oil immersion. The physical size of a camera pixel is approximately 11.5 pixels per \(\mu\)m.
Given that these datasets originate from different clinical sites, we refer to the respective clinical centers as C1, C2, C3, and C4. As mentioned in Sect. “Methodology”, for a domain incremental CL scenario, a fixed number of classes is required across all the episodes. We consider monocyte (MON), eosinophil (EOS), and lymphocyte (LYT) classes in each dataset. Classes other than MON, EOS, and LYT are excluded due to their limited availability in some datasets. More details regarding class selection are provided in the Supplements. Details regarding the number of samples, centers, and organs for each dataset are provided in Table 1. For an extensive evaluation, we create multiple CL experiments with the above-mentioned datasets. Specifically, we form four different sequences on these datasets, varying the dataset order in terms of increasing or decreasing dataset volume, as well as organ-specific learning. We mention the name of each of these four sequences, dataset order in each sequence, and the basis of ordering in Table 2.
Baselines
We compare our method against various CL benchmarks, including regularization-based approaches which do not keep memory buffers, but add a regularization term in the training loss, such as EWC11, SI12, and LwF13. Furthermore, we assess replay-based approaches that utilize buffers, including DER14, ER15, LR16, GEM17, and AGEM18. We also compare with GLRCL, a recent CL framework employing GMM-based generative replay for tumor classification in domain shift conditions6. Along with CL approaches, we also report the performances of non-CL methods, namely, the naive and joint strategy. In the naive approach, the model is fine-tuned using only the new task data without any mechanism to mitigate forgetting, thereby establishing a lower bound for the average performance across all tasks. In contrast, joint training assumes that all tasks are simultaneously available, enabling the model to train on all tasks concurrently and achieve an upper performance bound.
Classification models
Primarily, ImageNet-pretrained models like ResNet50 and its variants have been utilized for CL-based classification frameworks. However, the significant domain gap between natural and medical data may limit CL performance. To address this, we also validate for SSL-based models specifically tailored for pathology data. To this end, we used RetCCL19, a ResNet50 trained on a large cohort of human cancer pathology images. CTransPath20 and UNI21 are transformer based FMs with more capacity and trained on multi-scale with larger amount of data. DinoBloom22, an FM for hematology is not considered in our work as this backbone has been trained on variety of WBC datasets including the ones used in our experiments (Sect. “Datasets”). To construct our classification model, we select a backbone from {ResNet50, RetCCL, CTransPath, UNI}, discard the respective classification head and appended five learnable Fully Connected (FC) layers [512, 256, 128, 64, 32] for supervised classification. Each backbone is initialized with pre-trained weights from its respective repository. Following the approach in6, we freeze the backbone weights to preserve the KDE-based latent generator’s validity, training only the FC layers.
Training setup
We followed standard CL evaluation protocols using the Avalanche library23. The WBC images were resized to \(256 \times 256\) pixels, and batched with a size of 64. For our approach, we set best \(\alpha\) (regularization coefficient) by following grid search in set {0.01, 0.1, 0.2, 0.3, 0.4, 0.5} for each backbone. Similar to work6, we set the number of cluster centers for a domain to 10 in K-means clustering. Thus, the number of total cluster centers on which KDE is built is 40 latent vectors (\(10\times 4\) for 4 tasks in a sequence). For fair comparison with buffer based approaches we keep buffer size in GEM, AGEM, ER, LR, and DER as 40. We followed6 and set the number of components for per-domain GMM to 10. For SI, EWC, and LwF, the value of regularizing factor (\(\alpha\)) was set to 1 by following the literature6. All experiments were conducted using an NVIDIA A100 GPU, with training and evaluation sessions completing within a maximum of two hours across all methods, except for GEM which took a maximum of 15 hours. To ensure statistical reliability, all experiments were repeated five times, and we report the mean and standard deviation of the results.
Evaluation metrics
After sequentially learning \(T\) classification tasks, we constructed a train-test matrix2 \(P\), where each cell \(p_{ij}\) represents the accuracy on \(j^{\text {th}}\) dataset after completing training on \(i^{\text {th}}\) dataset. From this matrix, we computed CL metrics such as backward transfer (BWT)24, average accuracy after completing the last task (ACC)17, and Incremental Learning Metric (ILM)24, which measures the incremental learning capability of the approach. Higher values of these metrics indicate better performance. We detail the equations for these metrics in Table 3.
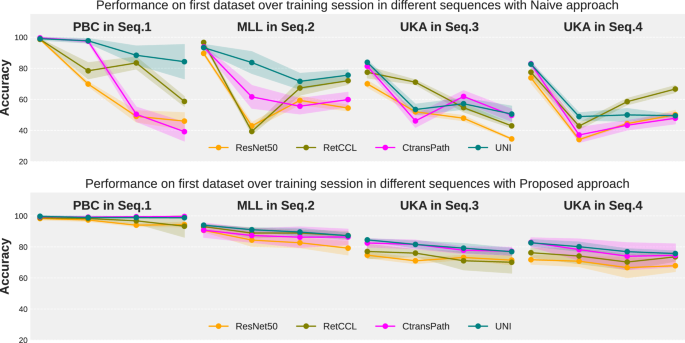
Performance of the first dataset upon learning other subsequent datasets in Seq.1 to Seq.4 in a naive (top row) and proposed (bottom row) way.
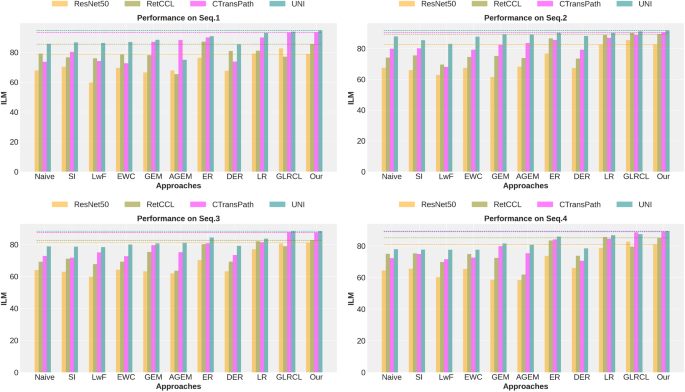
Backbone comparison in different approaches.
link








