An adaptive, continuous-learning framework for clinical decision-making from proteome-wide biofluid data
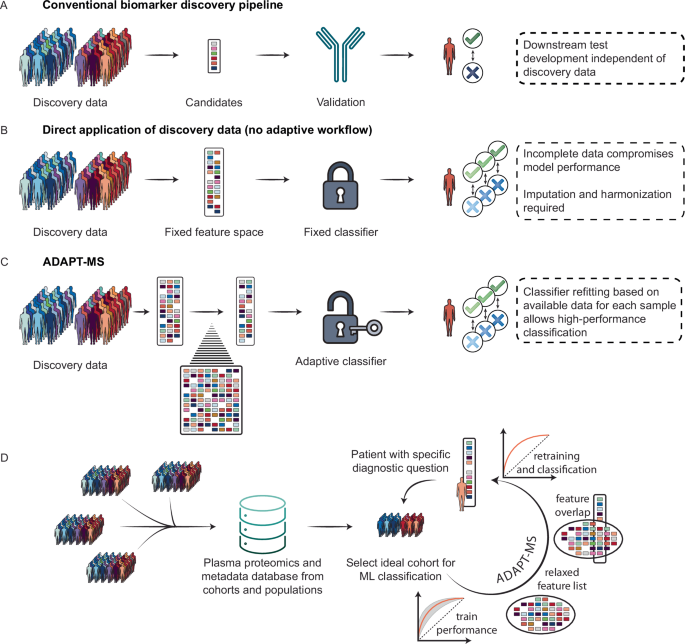
The ADAPT-MS framework
Current clinical proteomics pipelines rely on a rigid multi-step process: biomarker discovery is performed in a case–control cohort, followed by assay development for a fixed subset of proteins, and only then are these assays applied to classify new patient samples (Fig. 1A). This approach ensures data completeness through targeted measurements but limits flexibility, discards most of the proteome-scale data, and addresses only one diagnostic question at a time.
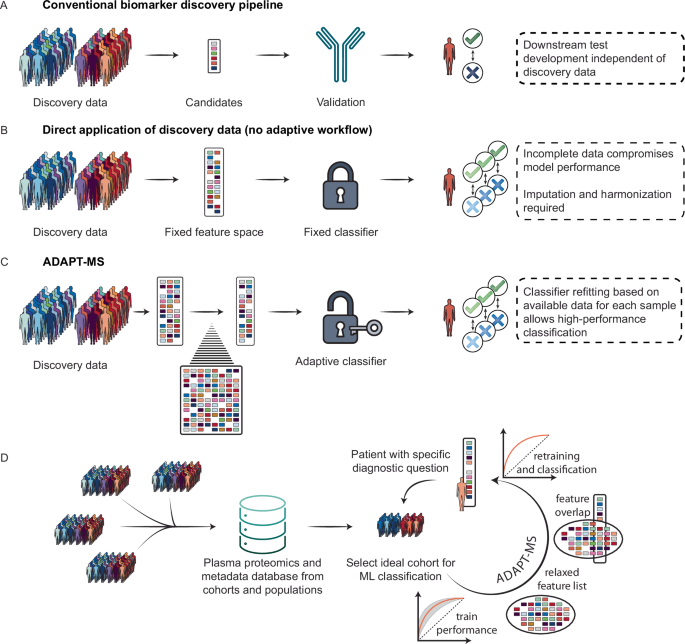
A In the conventional proteomics pipeline, candidate biomarkers are identified in a discovery cohort (e.g., case–control study), translated into targeted assays, and applied to new patients for a single diagnostic question. B Direct application of discovery proteomics data without adaptation to new patient samples suffers from incomplete data matrices, reducing classifier performance and often requiring imputation or harmonization. C ADAPT-MS overcomes this by dynamically retraining the classification model for each patient sample based only on the proteins actually measured, using a relaxed feature set derived from the training data. D For new diagnostic questions, ADAPT-MS retrospectively assembles matched training cohorts from large proteomics databases using sample metadata. Feature selection and classifier training are performed on this subset, enabling personalized, scalable, and continuous diagnostic applications based on unbiased proteome-wide measurements.
We developed a framework that enables direct use of discovery proteomics data for diagnostic classification, without the need for targeted assay development or imputation of missing values. The core innovation of our method—termed Adaptive Diagnostic Architecture for Personalized Testing by Mass Spectrometry (ADAPT-MS)—is the dynamic retraining of classification models on a per-sample basis. Rather than applying a fixed classifier across all samples, ADAPT-MS dynamically adapts the model to the proteins actually quantified in each individual sample (Fig. 1B, C). A relaxed feature list is first generated from the discovery dataset using cross-validation and feature selection. For each test sample, the overlap between its measured proteins and this feature list defines a custom feature set, on which a classifier is retrained using the original discovery data. The resulting model is used to assign a diagnostic label or score to that individual case.
This dynamic retraining strategy overcomes the long-standing problem of missing values which are always present in any discovery-mode proteomics. Subsequently, when applying a model on a single sample of interest, we only rely on features (proteins) that are actually detected. In this way, ADAPT-MS sidesteps the intricacies and potential biases of imputation or a simple zero-fill approach. It enables robust single-sample classification without requiring complete data matrices or bias-prone imputation, and allows direct diagnostic use of the proteomics measurement itself on single samples—transforming the output of a discovery experiment into a clinical decision tool.
Importantly, ADAPT-MS does not rely on a pre-defined case-control study for every diagnostic question. Instead, it leverages the increasing availability of large, population-scale proteomic datasets with linked clinical outcomes. For each new diagnostic task, the system can retrospectively assemble a suitable training cohort by identifying samples that match the patient of interest along relevant covariates, such as age, sex, comorbidities, and sampling context. This retrospective “cohort slicing” enables precise, hypothesis-specific modeling without requiring new prospective studies for every differential diagnosis. In effect, the framework replaces the traditional one-question-one-panel paradigm with a flexible, reusable infrastructure that draws diagnostic power from the full depth and scale of previously acquired proteome data (Fig. 1D). Illustrative clinical use cases for this flexible diagnostic approach are summarized in Table 1.
Dynamic retraining on plasma and serum proteomics cohorts
To evaluate the performance of the ADAPT-MS framework under real-world conditions, we applied it to a recently published dataset of plasma proteomes from patients assigned different subgroups of sepsis and healthy controls. This study, generated by the Roman Fisher group, represents one of the most comprehensive clinical proteomics efforts to date25. Among other sub-cohorts, this comprises a discovery cohort (902 samples) and a separate validation cohort (459 samples), all processed using high-throughput DDA in PASEF mode. We focused on a binary classification task: distinguishing sepsis patients with a specific transcriptomically defined sepsis response signature (SRS) from those without26,27. This biologically relevant subclassification is clinically challenging and had not been addressed in the original study. The data posed significant technical challenges typical for discovery proteomics, including variable and semi-stochastic missing values, especially for lower-abundance proteins (Supplementary Fig. 1A, B).
As a benchmark, we first trained a traditional machine learning model using the full discovery cohort: feature selection by random forest and classification with XGBoost. This pipeline achieved robust performance in cross-validation and yielded an area under the curve (AUC) of 0.83 on the independent validation set, demonstrating that proteomics data can support complex classification tasks when data completeness is ensured.
To apply the ADAPT-MS framework, we first extracted a relaxed set of protein features to support later sample-specific retraining. Relaxed in this context refers to features selected from randomly sampled cohort subsets in a cross-validation-like fashion, introducing controlled variability in the patient population and corresponding feature sets. This variability increases robustness in later patient-specific selection of features—by reducing dependence on any specific cohort composition. We performed a grid search over feature selection methods and classifiers (Supplementary Fig. 1C, D). As performance was consistent across most combinations (AUC 0.80–0.83), we selected t-test-based feature selection combined with logistic regression for its simplicity, robustness, and interpretability28. However, the ADAPT-MS framework is agnostic to both the feature selection method and classification algorithm employed. Classification tasks requiring non-linear decision-boundaries can readily incorporate algorithms, such as random forest or gradient boosting within the same ADAPT-MS architecture, simply by substituting the classifier while maintaining the sample-specific retraining strategy.
For each of the individual samples, we then identified the intersection between this relaxed feature list and the proteins actually quantified. A logistic regression model was then retrained on the discovery data using this intersection and used to classify the individual sample. This procedure was repeated independently for all validation samples. The ADAPT-MS model slightly outperformed the conventional fixed-classifier approach, reaching an AUC of 0.87 (Fig. 2A). Importantly, the performance remained stable even in samples with relatively high missingness, without requiring imputation. We further explored the number of features contributing to each prediction and found that performance plateaued at around 50 proteins per sample (Supplementary Fig. 1E). Correctly and incorrectly classified samples drew from similar distributions of available features over a broad range of requested features from feature selection, highlighting the method’s robustness to varying data completeness (Fig. 2B, Supplementary Fig. 1F). To further confirm the robustness of ADAPT-MS, we applied its retraining pipeline across different training set sizes and found that it consistently achieved high performance, comparable to conventional XGBoost classifiers on the same data (Fig. 2C). Computational time increased with training set size for both methods (Supplementary Fig. 1G, H). ADAPT-MS showed somewhat steeper growth during training, reflecting repeated feature-selection steps, but remained computationally lightweight overall (a one-time computational cost that takes only minutes on a standard laptop) and was faster than XGBoost during validation.
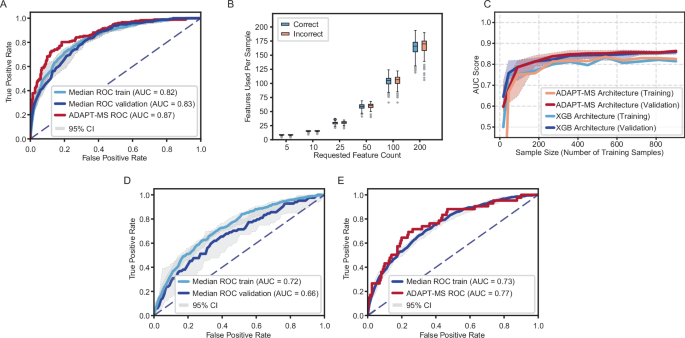
A Receiver Operating characteristic (ROC) analysis comparing ADAPT-MS to conventional fixed feature pipeline: ADAPT-MS outperforms a conventional fixed-feature pipeline (AUC 0.87 vs. 0.83). Shaded areas represent 95% confidence intervals from 10x repetition of classification for train and validation sets. The ADAPT-MS classifier is deterministic and has no defined confidence interval. B Robustness to missing values: correctly and incorrectly classified samples have similar numbers of features used, indicating consistent performance across varying degrees of missingness. Boxplots show median (line), interquartile range (box), and 1.5x interquartile range (whiskers) for the number of features used for classification per sample (461 samples classified in total in from the validation set). C Learning curves showing area under the ROC curve (AUC) as a function of training sample size for ADAPT-MS (Refit Architecture) and conventional XGBoost (XGB). Classification performance improves with increasing training set size for ADAPT-MS, highlighting its ability to benefit from continuous data accumulation. Shaded areas show 95% confidence interval from ten iterations. D RF – XGBoost architecture performance for the training and validation cohort sets of the MetS study. Gray areas show 95% confidence interval from ten iterations. E ADAPT-MS performance on the training cohort for the training architecture part and in single sample-retraining mode for the validation cohort. Gray areas show 95% confidence interval from ten iterations for the training data.
To illustrate the extension of ADAPT-MS to prognostic classification tasks, we applied our architecture to a recently published cohort investigating the risk prediction for development of metabolic syndrome (MetS) from serum proteomics29. Both discovery and validation cohorts were measured using unbiased discovery proteomics, making this dataset an ideal example case. The authors of the study applied a complex ML architecture wrapped around LightGBM gradient boosting; thus, it is not surprising that a standard RF—XGBoost model does not perform on par (Fig. 2D). However, the ADAPT-MS architecture yields equivalent performance to the published model (AUC 0.77) on the validation data, demonstrating the power of our simple and explainable yet readily applicable model (Fig. 2E).
These results demonstrate that ADAPT-MS can match or exceed standard machine learning pipelines while enabling true single-sample diagnostics and prognostics directly from discovery-mode proteomics. Our dynamic retraining strategy ensures that each classification is optimized for the data available in a given patient sample, making it suitable for routine clinical use even in the presence of variable missingness.
Generalizability of ADAPT-MS across clinical centers in CSF proteomics
To evaluate the robustness and cross-site generalizability of ADAPT-MS, we applied it to a multicenter cerebrospinal fluid (CSF) proteomics dataset of Alzheimer’s disease (AD) patients and controls. This dataset, generated in our own laboratory, comprises three independent clinical cohorts from Sweden, Berlin, and Magdeburg/Kiel, processed under comparable protocols but representing real-world variability in patient populations, pre-analytical bias and diagnostic accuracy30. The variability is visible by cohort-specific proteome signatures in unsupervised methods (Supplementary Fig. 2J) as well as in the different numbers of significantly changing proteins between AD patients and control samples across the different cohorts.
Each cohort included CSF proteomes acquired by data-independent acquisition (DIA) and annotated with standard AD biomarkers and clinical diagnoses. As previously reported, the Sweden and Magdeburg/Kiel cohorts showed clear separation between AD and non-AD patients, whereas the Berlin cohort exhibited substantial overlap, reflecting the challenge of cross-center reproducibility in clinical proteomics.
We first reanalyzed the raw data using an updated library-free DIA pipeline and reproduced the original classification performance using conventional machine learning (mean AUC 0.96, Supplementary Fig. 2A). Building on this, we applied ADAPT-MS using the Sweden cohort for discovery, with relaxed feature selection via cross-validation and t-tests (performance on 5x CV training cohort Sweden AUC 0.96, Supplementary Fig. 2B). We then dynamically retrained a logistic regression classifier for each individual test sample in the Berlin and Magdeburg/Kiel cohorts based on its quantified proteins and compared it to full dataset classification based on XGBoost following data frame imputation. ADAPT-MS achieved strong generalization performance across both external sites. When training on Sweden and testing on the other two cohorts, the AUC reached 0.85 (Magdeburg/Kiel) and 0.73 (Berlin)—despite the lower diagnostic signal in the latter cohort (Fig. 3A Supplementary Fig. 2C, D). Performance improved when training on a combined discovery cohort of Sweden and Magdeburg/Kiel, reaching an AUC of 0.80 in Berlin (Fig. 3A, Supplementary Fig. 2E, F). This demonstrates that ADAPT-MS benefits from diverse training data and scales well to complex, real-world settings. We also tested a multicentric diagnostic setup, where the training cohort was constructed from a balanced subset of patients from all three clinical sites. This strategy yielded strong and stable classification across cohorts, highlighting the flexibility of the framework to integrate heterogeneous data sources. Importantly, the method does not rely on full-cohort imputation or harmonization for any validation or actual application case on single samples, but adapts per sample based on the available feature space (Fig. 3B).
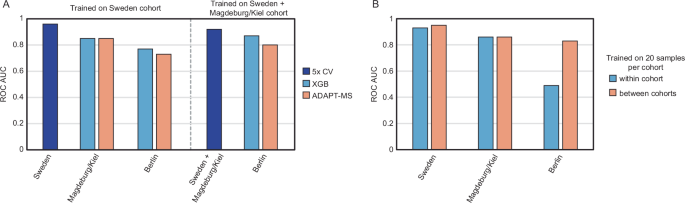
A Classification performance (ROC AUC) of ADAPT-MS compared to fixed-feature and XGBoost classifiers in two cross-center settings: training on Sweden alone, and training on Sweden combined with Magdeburg/Kiel, both tested on Berlin. ADAPT-MS outperforms or matches alternatives across scenarios. B Performance of ADAPT-MS using only 20 training samples per cohort, comparing within-cohort and cross-cohort classification. ADAPT-MS achieves consistently high AUCs even in cross-site settings, highlighting its adaptability across heterogeneous datasets without the need for imputation or panel harmonization.
Together, these results underscore a key advantage of ADAPT-MS: the ability to support individualized diagnostics across clinical sites, even in the presence of technical and biological variation. This cross-center generalizability is essential for real-world deployment of proteomics-based diagnostics.
Retrospective cohort matching improves classification performance in simulated diagnostics
To further evaluate the retrospective cohort selection component of the ADAPT-MS framework, we again turned to the sepsis proteomics dataset used above. While the SRS signature has been defined by transcriptomics to classify patients with different forms of sepsis and proteomics is able to predict those classes well (Fig. 2), Mi et al. defined a purely proteome-based classification of sepsis patients based on unsupervised proteome clustering (SPC1/2/3)25. This illustrates the advantage of ADAPT-MS, being able to adaptively and retrospectively slice and match cohorts for diagnostic decisions. Based on the proteomics data, any newly acquired patient sample can be classified into i) sepsis or non-sepsis, based on a classifier built on the cases vs control samples of the cohort/database, ii) SRS or non-SRS type of sepsis and iii) into the newly defined proteomics sepsis classes (SPC1/2/3) from Mi et al. (Fig. 4A–C). Further, any other covariate present with sufficient biological effect size could be added to the possible classifications. For example, if the control samples would include a large number of patients with acute inflammation but not sepsis, this would be possible to differentiate and diagnose with a separate classifier.
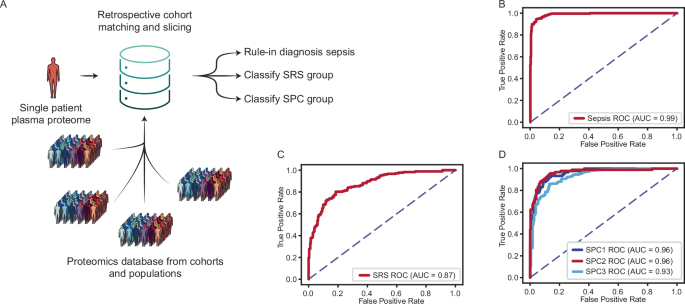
A Schematic of the ADAPT-MS workflow for retrospective cohort matching. A single patient’s proteome-wide measurement is reused to address multiple diagnostic questions. For each task, a matched training cohort is dynamically selected from a reference database using metadata (e.g., age, sex, comorbidities), and a sample-specific classifier is retrained. B Classification performance of ADAPT-MS for sepsis vs controls (AUC = 0.99). C Classification performance of ADAPT-MS for SRS vs non-SRS (AUC 0.87). D Classification performance of ADAPT-MS for sepsis plasma proteome-based clusters (SPC1/2/3), with AUCs ranging from 0.93 to 0.96.
This highlights a key strength of ADAPT-MS: tailoring diagnostics to individual cases using retrospective matching. In real-world scenarios where diseases are heterogeneous and prospective control cohorts are impractical, this approach would enable flexible, scalable diagnostics grounded in existing data. It also addresses the n × m scaling problem in diagnostic development by reusing the same sample against dynamically assembled reference groups—supporting multiple hypotheses without remeasurement.
Crucially, these gains come with no increase in computational or practical complexity. Each retraining step takes only a few seconds, even on standard hardware, and uses simple, well-established statistical methods, such as ANOVA for feature selection and logistic regression for classification. This deliberate simplicity promotes interpretability and generalizability, avoiding the overfitting often associated with complex, highly parameterized machine learning or deep-learning models. By design, the system is transparent and stable, favoring robustness over marginal improvements in performance.
The flexibility of retrospective cohort matching further enhances diagnostic resolution. Depending on the clinical context, the reference population can be broadly defined—e.g., healthy individuals of the same age and sex—or finely tuned to match the characteristics of the patient under consideration. This includes the possibility of constructing a near-identical comparator group, effectively creating a “digital twin” for highly personalized diagnostics. Because the full proteomic profile is measured once and retained, each sample can be interpreted repeatedly and also longitudinally, against different reference groups and for multiple diagnostic hypotheses, without remeasurement or assay development.
link








