A hybrid vision transformer with ensemble CNN framework for cervical cancer diagnosis | BMC Medical Informatics and Decision Making

In this section, the description of the developed models for detecting cervical cancer by utilizing Pap-smear images is provided by applying transfer learning with ensemble learning and ViT transformer techniques. Initially, cervical cell cancer data from the Mendeley LBC and SIPaKMeD datasets, which contain Pap smear images, were prepared. Figure 1 illustrates the sequential procedure of constructing the model, beginning with the splitting of the data and the augmentation of the training dataset. Each dataset is individually used to develop models, and then the two datasets are combined to train models again, including a variety of equipment and tools to capture cervical cells. Subsequently, the pre-trained transfer learning models are introduced individually. Afterward, the high-level features are combined to form the ensemble learning model, which acts as the fundamental framework for the convolutional network. In the end, the model is fed into the transformer encoder model to classify cervical cell cancer, while also offering an explainable analysis.
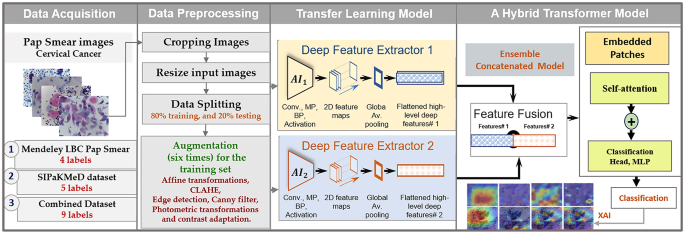
An overview of the proposed framework designed to classify the multi-class cervical cancer cells using the combined datasets
Dataset description
Mendeley LBC pap smear slides dataset
Mendeley LBC Pap Smear was obtained from Babina Diagnostic Pvt., India [40]. This data consisted of information from 460 patients who had undergone cervical screening examinations. It has four categories of cervical cancer: Negative, LSIL (Low-grade intraepithelial lesions), HSIL (High-grade intraepithelial lesions), and SCC (Squamous Cell Carcinoma). There was a total of 963 images that were taken from Pap smear slides with a magnification of 400 times. Of these, 613 images were classified as belonging to the normal or NILM group, while 350 images were classified as belonging to the abnormal category.
SIPaKMeD dataset
During the year 2018, academics affiliated with the University of Ioannina in Greece were responsible for the production of the dataset [27]. In the SIPaKMeD database, there are 4049 images of women’s cells that have been meticulously isolated from 966 cluster cell images of Pap smear slides. These images may also be added to the database. An OLYMPUS BX53F optical microscope was used to acquire the images, which were obtained through the utilization of a CCD digicam modification. Sup-intermediate, Parabasal, Koilocytotic, Dyskeratotic, and Metaplastic cancer cells are included in the dataset, among the majority of cancer cells that have been classified into these five groups. Table 2 shows the description for both datasets, including URL access, type of images, and categories.
Data preprocessing, splitting, and augmentation
For preprocessing, choosing the region of interest (ROI) with cervical cancer cells and disregarding the rest of the image-produced patch images. A better method for boosting extremely dark images using histogram equalization, such as contrast-limited adaptive histogram equalization (CLAHE). The image patches are rescaled to 256 × 256 pixels in size. Both datasets, Mendeley LBC and SIPaKMeD, are divided into training, validation, and testing sets, with each class receiving 80%, 10%, and 10% of the total data, respectively, as shown in Tables 3 and 4. Image processing and geometric transformations, such as those obtained from the “imgaug” library, are utilized to carry out data augmentation [31]. To increase the number of training samples, the augmented images are produced alongside the original training images. The augmentation procedure is repeated six times during the process using the affine transformations, CLAHE, edge detection, Canny filter, photometric transformations, and contrast adaptation by ImageDataGenerator [41] to images that have been altered.
Once the two datasets have been merged, the training set, validation set, and testing set will contain a total of 3484, 772, and 755 images, respectively. These images are spread across 9 different classes.
AI models
DenseNet201
The DenseNet201 represents an advanced iteration derived from the dense-network framework. This architectural design employs a unique approach by creating direct connections between every layer in a feed-forward manner, facilitating interactions among all constituent layers [42]. Moreover, the DenseNet201 architecture incorporates multiple pooling layers while maintaining a compact structural form. Consequently, these design choices contribute to a reduction in both the number of parameters and the overall complexity of the model, thereby improving its operational efficiency.
InceptionResNetV2
The stem of the network is based on InceptionV4; however, the architecture of the network is comparable to that of InceptionResNetV1 [43]. There is a shortcut link located on the far-left side of each module. A combination of inception architecture and residual connections is used to achieve superior classification results. Both the input and the output of the convolutional operation in the inception module must be identical for the inception convolutional operation to become successful.
Xception
The Xception model, a type of deep convolutional neural network, has demonstrated significant advancements in its use for medical classification tasks. The pre-trained version of the network is 71 layers deep and has been trained on over a million images from the ImageNet database [44]. The development of this technology was undertaken by researchers employed by Google. Google introduced the concept of Inception modules in convolutional neural networks as a transitional stage between normal convolution and the depth-wise separable convolution operation. The layers below 106 have been fine-tuned and made untrainable.
The proposed AI ensemble and transformer encoder learning models
Applying ensemble learning principles has led to the development of various DL models instead of relying solely on an individual DL model [45]. Consequently, integrating these multiple models represents the final phase in the prediction process [46]. Enhanced classification outcomes can be achieved by leveraging a broader range of pertinent data from differing classifiers. The ensemble method for feature extraction introduced in this study integrates two pre-trained DenseNet101 and InceptionResNetV2 deep learning models, as illustrated in Fig. 2.
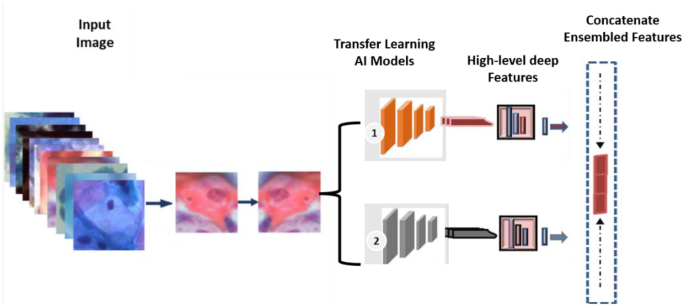
The architecture of the proposed concatenated ensemble-based features model
In the proposed ensemble learning framework, deep features are extracted from two CNN architectures: DenseNet101 and InceptionResNetV2. To obtain these features, the classification layers of each model are removed, and the final convolutional blocks are utilized for deep feature extraction. The hybrid model is constructed by fusing the high-level representations obtained from both CNN backbones. Specifically, global average pooling (GAP) is applied to the output of each model to transform the extracted feature maps into vector representations. These high-level feature vectors are then concatenated to form a comprehensive feature space. The resulting fused feature vector is subsequently embedded and passed to the prediction stage, which is based on a ViT architecture.
A transformer is a type of DL model that uses a self-attention mechanism to give different parts of the input data different levels of importance. It does this by using an encoder-decoder architecture. This study employs the ViT framework, integrating an encoder methodology to identify early-stage cervical cancer from pap-smear images. ViT encoder yields two outputs from the processed input image: a comprehensive token embedding vector that spans all layers, as well as the attention weights from each head. The self-attention mechanism facilitates the production of a representation for a singular input sequence by creating relational connections among various positions within that sequence [47, 48]. Each block integrates a normalization layer alongside residual connections. The encoded framework is composed of a self-attention mechanism and an MLP component. Each block integrates a normalization layer alongside residual connections. The multilayer perceptron is a specific kind of feed-forward neural network that features dense layers and dropout layers [49] as outlined in the equations:
$$\:\:Attention\left(Q,K,V\right)=Softmax\left(\frac{{Qk}^{t}}{\sqrt{{d}_{k}}}\right)v,$$
(1)
where Q means query vector, V is a value-dimensional vector, and K refers to the key vector. The \(\:{d}_{k}\) represents the variance of the product \(\:Q{k}^{t}\), which has a zero mean. In addition, normalizing the product by dividing it by the standard deviation \(\:\sqrt{{d}_{k}}\). The SoftMax function transforms the scaled dot product into attention scores, which are central to the transformer’s ability to attend to various parts of the input image in parallel. This mechanism enables the model to understand the image content comprehensively by simultaneously focusing on multiple aspects of the input. The multi-head attention mechanism enhances this capability by projecting the queries, keys, and values into multiple subspaces through learned linear transformations [50]. These projections are performed h times, allowing the model to attend to information from different representation subspaces concurrently. The computation of multi-head attention is formally defined in Eqs. (2,3).
$$\:MultiHead\left(Q,K,V\right)=Concat\left({head}_{i},\dots\:\dots\:.{head}_{h}\right){W}^{o\:}$$
(2)
$$\:{Head}_{i}=\text{A}\text{t}\text{t}\text{e}\text{n}\text{t}\text{i}\text{o}\text{n}(\text{Q}{\text{W}}_{i}^{Q},\:\text{K}{\text{W}}_{i}^{K},\text{V}{\text{W}}_{i}^{V})$$
(3)
where the projections are parameter matrices \(\:{\text{W}}_{i}^{Q}\in\:{\text{R}}^{{d}_{model}\:x\:{d}_{k}},\:{\text{W}}_{i}^{K}\in\:{\text{R}}^{{d}_{model}\:x\:{d}_{k}}\), \(\:{\text{W}}_{i}^{V}\in\:{\text{R}}^{{d}_{model}\:x\:{d}_{v}}\) and \(\:{W}^{o}\in\:{\text{R}}^{{hd}_{v}\:x\:{d}_{model}}\). On the other hand, the MLP block consisted of a non-linear layer of Gaussian error linear unit (GELU) with 1024 neurons and batch normalization, with a dropping rate of 50% across all dropout layers. Attention.
Explainable AI
Explainable AI (XAI) is a growing area of study in machine learning, particularly in the field of medicine. A philosophy of explainable artificial intelligence for static images is that a high-performing DL model accurately recognizes the relevant part of the image without placing excessive emphasis on insignificant details [51]. There are multiple approaches for creating visual representations of predictions from image classification models, many of which utilize post hoc techniques. These techniques involve examining the output of a DL model by employing various algorithmic tools to generate insightful explanations [52]. The ReLU function was employed to mitigate the influence of negative weights on a particular class. This approach operates under the assumption that the spatial elements within the feature maps associated with negative weights are likely representative of distinct categories present in the image [53], as illustrated by Eq. (4).
$$\:Grad\_{M}_{c}(x,y)=ReLU(\:{\sum\:}_{k}{\alpha\:}_{k}^{c}{f}_{k}\left(x,y\right)$$
(4)
Where \(\:{f}_{k}\left(x,y\right)\) denotes the activation at the spatial element (x, y) in the kth feature map, and the weight \(\:{\alpha\:}_{k}^{c}\) is calculated by finding the gradient of a prediction score, 𝑆𝑐, for the k-th feature map.
$$\:{\alpha\:}_{k}^{c}=\:{\sum\:}_{x,y}\frac{{\delta\:S}_{c}}{{\delta\:f}_{k}(x,y)}$$
(5)
Experimental setup
The proposed AI hybrid model is characterized by its end-to-end training approach. For both our training phase and the early stopping callback, a learning rate of 0.001 was implemented, utilizing an Adam optimizer with a clipping value of 0.2 and a patience parameter set to 30. Our training methodology for all AI models comprises a comprehensive iteration over 100 epochs, aimed at fine-tuning the hyperparameters effectively. The image is 256, a patch size of 2, and an input dimension of 20, complemented by a drop rate of 0.01 applied across all layers, along with 8 attention heads. In this context, to illustrate the transformation of high-dimensional vectors into lower-dimensional counterparts while ensuring no loss occurs, the parameters for both embed_dim and num_mlp have been meticulously calibrated through empirical techniques, achieving optimized values of 64 and 256, respectively.
Evaluation strategy
The performance of the proposed model in classification tasks was evaluated using the metrics accuracy, area under the receiver operating characteristic curve (AUC-ROC), sensitivity (recall), specificity, and F1-score, that used as in our previous work [46].
Accuracy quantifies the proportion of correct predictions relative to the total number of cases. It is defined as:
$$\:Accuracy\:\left(ACC\right)=\frac{TP+TN}{TP+FP+TN+FN}$$
(6)
Here, TP, TN, FP, and FN represent true positives, true negatives, false positives, and false negatives, respectively. Accuracy serves as an overall measure of the model’s performance, but may be less informative in cases of class imbalance.
The recall represents the proportion of actual positives that were correctly identified by the model. It is computed as:
$$\:Recall\:\left(REC\right)=\frac{TP}{TP+FN}$$
(7)
Precision indicates the proportion of true positive predictions among all positive predictions, calculated as:
$$\:Precision\:\left(PRE\right)=\frac{TP}{TP+FP}$$
(8)
F1-score provides a balanced measure of precision and recall by calculating their harmonic mean. It is defined as:
$$\:F1-score=2.\:\frac{Precision\:.\:Recall}{Precision+Recall}$$
(9)
A score of 1 indicates perfect performance in both precision and recall, while a score of 0 reflects the poorest performance. This metric is especially useful in scenarios with imbalanced datasets, as it considers both false positives and false negatives.
The ROC curve plots true-positive rate vs. false-positive rate over every decision threshold. The AUC condenses this plot into a single score from 0 to 1: closer to 1 means the classifier separates the two classes well; 0.5 is random guessing. Together with a confusion matrix showing counts of TP, FP, TN, and FN, you can compute accuracy, sensitivity, specificity, and F1-score, giving a rounded picture of performance, especially useful for imbalanced, binary pathology-classification problems [54].
Execution environment
Experiments were run on an MSI GS66 laptop (Windows 11) with an 11th‑gen Intel Core i7‑11,800 H CPU, 32 GB RAM, a 2 TB NVMe SSD, and an RTX 3080 GPU (16 GB RAM). The code was written in Python 3.10 using Keras with a TensorFlow backend.
link








