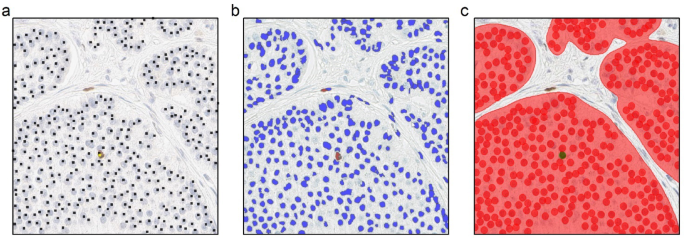
Achieving automatic tumor cell detection and Ki67 score estimation is one of the central goals for the application of image analysis in digital pathology. While numerous studies have already addressed the development and evaluation of related image analysis tools32 many of these studies focused solely on the number of detected cells or the Ki67 score33. However, even if an image analysis tool returns the correct number of detected cells, the same as the reference standard, it does not guarantee that the correct cells have been identified (Supplementary Table S3 and S4 for detailed counts of tumor cells). The current study aims to evaluate detection results at the individual cell level to fully understand any performance differences. To achieve this, performance of a ML-based and a non-ML-based image analysis tool were evaluated in estimating Ki67 score in NETs. The employed non-ML analysis tool represents an example for the conventional approach with a user-friendly interface. While the chosen ML-based platform, supported by strong customer service, exemplifies the state-of-the-art technology and facilitates easy implementation for first-time users. By emphasizing the importance of correct tumor cell identification, our study provides a more comprehensive and clinically relevant evaluation of image analysis tools, ultimately enhancing their trustworthiness and interpretability.
This study specifically focused on challenging NET cases with a low Ki67 score of less than 5%, as accurate estimation of Ki67 score is particularly important for these low proliferating tumors, where a prognostically significant cut-off value of 3% separates G1 from G2 tumors32. Misclassifying these cases can have significant therapeutic implication. The WHO recommendation to count 500-2,000 cells is rather vague and does not consider the dependency on the proliferation rate. However, studies in breast cancer have clearly demonstrated that an accurate result depends on both the proliferation rate and the number of cells evaluated4. Since manually counting such a large number of cells is impractical, an automated method that can assist in differentiating between G1 and G2 NETs can significantly improve workflow efficiency.
This study demonstrated that neither of the two employed image analysis tools displayed perfect performance after comparing their result at the cell level with the reference standard. However, the ML-based analysis with Aiforia exhibited better agreement with the reference standard in both cell detection and Ki67 score calculation compared to the non-ML-based analysis with ImageScope. Furthermore, it was demonstrated that the improved concordance between the Aiforia’s Ki67 score, and the reference standard was attributed to its ability to distinguish between tumor and non-tumor cells.
The superior performance of the ML-based tool may stem from its use of supervised learning, which enables it to adapt to the data and learn parameters instead of relying on fixed, predefined parameters34. However, the performance of Aiforia also showed limitations, manifested by its tendency to pick more positively Ki67-stained cells, including (i) faintly stained Ki67 positive tumor cells which were dismissed by the pathologist (SL), or (ii) Ki67 positively stained proliferating non-tumor cells, resulting in a slightly higher estimated score.
The issue with the detection of faintly stained cells can be attributed to the general difficulty in visually assigning correct labels for weakly stained tumor cells. For those cells, labeling might be highly subjective due to stain quality, section thickness, nucleus appearance, as well as inter-observer and intra-observer variability35. The issue of non-tumor cell detection could be due to several underlying reasons, such as limited variation in the training dataset, and narrow control over the training procedures and other hyperparameters, which may limit the ability to fine-tune the model. Vesterinen et al.. reported a mildly better agreement between their trained Aiforia model and manual counting33. This could be due to having a bigger dataset. Another possible factor could be differences in the neural network settings. Additionally, they reported only the Ki67 score and did not provide details about their model’s performance in terms of cell counts.
Two main shortcomings in the performance of the non-ML analysis with ImageScope were observed, namely (i) difficulties distinguishing tumor from non-tumor cells, and (ii) issues related to the segmentation of individual cells. Specifically, in the presence of lymphocytes, endothelial cells, and stromal cells, the algorithm would frequently have difficulty distinguishing tumor cells (Supplementary Fig. S5). Depending on the distribution of cell types within a sample, this issue would frequently lead to the over-counting of tumor cells (Supplementary Table S3 and S4 for detailed counts of tumor cells). Non-ML-based analysis tools typically operate with a set of predefined parameters, and thus lack flexibility and adaptability when dealing with complex tasks. One possible solution to address this issue is to limit the scope of analysis, for example to analyze only tumor areas. Using a similar method, Volynskaya et al.. reported that their ImageScope analysis correlated with the pathologists’ results, after manual removal of the stroma within the analyzed regions36. While such an approach can improve the analysis results by excluding most non-tumor cells from the analysis, it is more time-consuming and requires domain expertise for precise annotation, making it impractical in our opinion. Other researchers have attempted different strategies, such as (virtual) double staining32 or finding complementary biomarkers37,38,39 to facilitate distinction between tumor and non-tumor cells.
The second shortcoming in the performance of ImageScope was related to imprecise segmentations. Two main issues were observed in this regard. First, the method often struggled to accurately distinguish overlapping tumor cells, leading to an underestimation of the number of individual tumor cells (Supplementary Fig. S2). Secondly, ImageScope faces challenges in correctly segmenting Ki67 positive tumor cells in cases of inhomogeneous staining. This issue resulted in overestimation, where one cell was segmented as multiple objects (Supplementary Fig. S3). One approach to overcome the segmentation problem is to measure the area fraction, which compares the area of brown-stained nuclei to the total area of brown- and blue-stained nuclei40. However, this solution can be sensitive to tumor type and cell size, making it less universally applicable. While efforts have been made to address this issue21 a universally applicable solution has not yet emerged.
Our study stands out as one of the few that specifically investigated the use of ML for estimating the Ki67 score in gastroenteropancreatic NETs. Other similar studies have also developed ML models, either using proprietary software33 or open source software32,41,42. Using proprietary software, like Aiforia43,44 offers certain advantages, such as fast and reliable customer support and often a more intuitive user interface. These factors make it easier, even for pathologists, to fine-tune an ML model. However, there are potential issues associated with proprietary closed source software, including cost, limited insight and control over individual settings and parameters, and ownership of the final trained model. Conducting similar analyses using open source software provides an alternative approach. In this case, more technical knowledge is required to implement and fine-tune the model, but it avoids the issues associated with proprietary software. The choice between proprietary software and open source software depends on various factors, including the specific requirements of the study, available resources, and the technical expertise of the researchers.
Even though Ki67 staining is a routine procedure in most histology laboratories, there can be a substantial amount of staining variation in terms of the brown color of positively Ki67-stained cells. This variation has the potential to affect prediction results across different slides. While the current study takes into account the presence of stain variations when tuning algorithms, additional approaches specifically targeting this issue could involve the use of stain normalization techniques45or virtual Ki67 staining methods46. An even more effective strategy could be the application of virtual double Ki67/synaptophysin staining, which not only addresses staining variation, but also helps in identifying tumor cells, facilitating a better distinction between tumor and non-tumor cells47.
A common challenge in evaluating automated tumor cell detection tools is the lack of an absolute ground truth for comparison. Instead, an approximative ground truth is usually established by a domain expert. In our study, we considered the manual counting of tumor cells by an experienced gastrointestinal pathologist to be a fair representation of the ground truth. The choice of using only one domain expert in setting the reference standard, i.e., not considering the expected inter-observer variability, could be regarded as an additional limitation. However, we believe this approach is reasonable considering the focus of the study, which is comparing the performance of two image analysis tools in detecting the correct tumor cells.
While hotspot selection is often considered a central step in Ki67 scoring, there are no precise instructions on how to choose a hotspot or assess one; for a discussion of this topic see for example Volynskaya et al.36. Some studies have explored the reliability of Ki67 scores when averaging several smaller hotspots or using a single larger hotspot48,49,50,51while others have employed automatic hotspots selection52,53,54. Since the Ki67 score is affected both by the number and size of chosen hotspots, any differences in these parameters between the compared methods could make direct comparison unreliable. In light of these issues, we chose to use the term ROI rather than the term hotspot and decided to define ROIs with a standard size containing approximately 200 cells, which was considered adequate to meet the aim of this study – to compare performance’s accuracy of two image analysis tools in cell level.
In conclusion, accurately determining the Ki67 score is crucial for grading NETs, predicting patient prognosis, and guiding treatment decisions. The integration of automated image analysis tools in clinical practice can enable pathologists to evaluate larger portion of the biopsy in less time. In the current study, we compared two image analysis tools for cell detection and Ki67 score estimation in NETs with low proliferation index and highlighted the importance of evaluating performance at the individual cell level. In comparison with the chosen reference standard, the ML based image analysis demonstrated a better agreement in correctly detecting individual tumor cells, both Ki67 positive and negative, compared to the non-ML approach, resulting in improved Ki67 score calculation.
Our study is among the few publications that compares image analysis tools in a diagnostic setting at the individual cell level. Unfortunately, there is a lack of comprehensive evaluations and comparisons of image analysis tools, particularly in the rapidly expanding market of commercially available artificial intelligence software. The lack of such evaluations hampers the ability of pathology departments to make informed decisions when purchasing these tools. Additionally, existing publications often provide incomplete assessments of model performance, further highlighting the need for more rigorous and comprehensive evaluations.
In the future, we aim to integrate the ML-based image analysis tool in routine diagnostics. A possible integration scenario can be for ML to flag the borderline cases, those near the G1/G2 threshold, for a more comprehensive manual review. This would help reduce the consequences of systemic bias and misclassification by image analysis tools. It can also output confidence scores, enabling pathologists to focus on low confidence predictions. This will involve further validation studies, including detailed analyses of comprehensive datasets of NET subtypes, outcome studies in patients with NETs, as well as the use of large external tumor datasets to validate the performance of the image analysis tool. Clinical validation, which involves correlating the calculated Ki67 score with clinical outcome data, is another important step to ensure the reliability and usefulness of the algorithm in real-world scenarios.
link